Any Sklearn Regressor as a Survival Model — Does It Actually Work? Benchmarking vs Established Packages
This study compares (link to notebook at the end of this post) survivalist — a model-agnostic (allows you to plug in any scikit-learn-like regressor as the base learner), probabilistic survival analysis package — against established survival analysis packages (scikit-survival, lifelines) on 15 real-world datasets spanning oncology, HIV/AIDS, genomics, criminology, transplant medicine, cardiac surgery, and political science.
Key contributions of this study:
survivalistis benchmarked with every scikit-learn regressor (default hyperparameters, defensive programming)- Metrics: C-index, Integrated Brier Score (IBS), Brier Score at fixed time horizons
- Datasets: 15 benchmark survival datasets (see Section 2)
- Baselines:
lifelinesCoxPH,scikit-survivalCoxPH + Random Survival Forest + Gradient Boosting Survival
Author note: All
survivalistmodels useSurvivalCustomwrapping sklearn regressors (default hyperparams).
Failures are caught defensively and logged — no experiment crashes the whole study.
Dataset Overview
| Dataset | Source | n | p | Event rate | Domain |
|---|---|---|---|---|---|
| WHAS500 | survivalist | 500 | 14 | ~0.42 | Cardiac (heart attack) |
| GBSG2 | survivalist | 686 | 8 | ~0.44 | Breast cancer |
| VeteransLungCancer | survivalist | 137 | 6 | ~0.93 | Oncology |
| FLChain | sksurv | 7,874 | 9 | 0.28 | Clinical (serum proteins) |
| AIDS | sksurv | 1,151 | 11 | 0.08 | HIV/AIDS trial |
| BreastCancerGenomic | sksurv | 198 | 80 | 0.26 | Genomics (high-dim) |
| Rossi | lifelines | 432 | 7 | 0.26 | Criminology (recidivism) |
| KidneyTransplant | lifelines | 863 | 4 | 0.16 | Transplant medicine |
| NCTCGLung | lifelines | 167 | 8 | 0.28 | Oncology (NCCTG) |
| LymphNode | lifelines | 686 | 8 | 0.25 | Breast cancer (lymph) |
| Leukemia | lifelines | 42 | 3 | 0.71 | Oncology (small n) |
| Larynx | lifelines | 90 | 4 | 0.56 | ENT oncology |
| Lymphoma | lifelines | 80 | 1 | 0.68 | Oncology (lymphoma) |
| StanfordHeart | statsmodels | 69 | 1 | 0.65 | Cardiac surgery |
| DemocracyDuration | lifelines | 1,808 | 9 | — | Political science |
# ── install all required packages ────────────────────────────────────────────
# Uncomment as needed
!pip install survivalist --upgrade
!pip install scikit-survival
!pip install lifelines
"""## 1. Imports & Global Config"""
import warnings
import traceback
import time
import inspect
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from tqdm.auto import tqdm
warnings.filterwarnings("ignore")
RANDOM_STATE = 42
TEST_SIZE = 0.25
N_SPLITS_CV = 3 # for cross-val inside baseline models
sns.set_theme(style="whitegrid", palette="muted")
plt.rcParams["figure.dpi"] = 120
print("✓ base imports ok")
"""## 2. Dataset Loaders
We load 15 survival datasets from `survivalist.datasets`, `scikit-survival`, `lifelines`, and `statsmodels`.
All datasets are preprocessed uniformly: one-hot encoding, median imputation, structured array output.
"""
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
# ── survivalist datasets ──────────────────────────────────────────────────────
from survivalist.datasets import (
load_whas500,
load_gbsg2,
load_veterans_lung_cancer,
)
# ── scikit-survival extra datasets ───────────────────────────────────────────
from sksurv.datasets import (
load_flchain,
load_aids as sksurv_load_aids,
load_breast_cancer as sksurv_load_breast_cancer,
)
# ── lifelines datasets ────────────────────────────────────────────────────────
from lifelines import datasets as ll_datasets
# ── statsmodels ───────────────────────────────────────────────────────────────
import statsmodels.datasets.heart as sm_heart
def _encode(df: pd.DataFrame) -> pd.DataFrame:
"""One-hot encode object/category columns; convert bools to int."""
cat_cols = df.select_dtypes(include=["object", "category"]).columns.tolist()
df = pd.get_dummies(df, columns=cat_cols)
bool_cols = df.select_dtypes(include=["bool"]).columns.tolist()
df[bool_cols] = df[bool_cols].astype(int)
return df
def _clean(X: pd.DataFrame) -> pd.DataFrame:
"""Fill NaNs, drop zero-variance columns."""
X = _encode(X.copy())
X = X.fillna(X.median(numeric_only=True))
X = X[[c for c in X.columns if X[c].nunique() > 1]]
return X
def _make_y(events, times):
"""Build sksurv-compatible structured array."""
return np.array(
[(bool(e), float(t)) for e, t in zip(events, times)],
dtype=[("event", bool), ("time", float)],
)
def _split(X, y):
return train_test_split(X, y, test_size=TEST_SIZE, random_state=RANDOM_STATE)
# ────────────────────────────────────────────────────────────────────────────
# Individual loaders
# ────────────────────────────────────────────────────────────────────────────
def _load_whas500():
X, y = load_whas500()
return _clean(X), y
def _load_gbsg2():
X, y = load_gbsg2()
return _clean(X), y
def _load_veterans():
X, y = load_veterans_lung_cancer()
return _clean(X), y
def _load_flchain():
X, y = load_flchain()
events = [r[0] for r in y]
times = [r[1] for r in y]
return _clean(X), _make_y(events, times)
def _load_aids():
X, y = sksurv_load_aids()
events = [r[0] for r in y]
times = [r[1] for r in y]
return _clean(X), _make_y(events, times)
def _load_breast_cancer_genomic():
X, y = sksurv_load_breast_cancer()
events = [r[0] for r in y]
times = [r[1] for r in y]
return _clean(X), _make_y(events, times)
def _load_rossi():
df = ll_datasets.load_rossi()
X = _clean(df.drop(columns=["week", "arrest"]))
y = _make_y(df["arrest"], df["week"])
return X, y
def _load_kidney_transplant():
df = ll_datasets.load_kidney_transplant()
X = _clean(df.drop(columns=["time", "death"]))
y = _make_y(df["death"], df["time"])
return X, y
def _load_ncctg_lung():
df = ll_datasets.load_lung().dropna()
events = (df["status"] - 1).astype(int) # 1=censored→0, 2=dead→1
X = _clean(df.drop(columns=["time", "status"]))
y = _make_y(events, df["time"])
return X, y
def _load_lymph_node():
df = ll_datasets.load_lymph_node()
drop = ["rectime", "censrec", "survtime", "censdead",
"diagdateb", "recdate", "deathdate"]
X = _clean(df.drop(columns=drop))
y = _make_y(df["censdead"], df["survtime"])
return X, y
def _load_leukemia():
df = ll_datasets.load_leukemia()
X = _clean(df.drop(columns=["t", "status"]))
y = _make_y(df["status"], df["t"])
return X, y
def _load_larynx():
df = ll_datasets.load_larynx()
X = _clean(df.drop(columns=["time", "death"]))
y = _make_y(df["death"], df["time"])
return X, y
def _load_lymphoma():
df = ll_datasets.load_lymphoma()
X = _clean(df.drop(columns=["Time", "Censor"]))
y = _make_y(df["Censor"], df["Time"])
return X, y
def _load_stanford_heart():
df = sm_heart.load_pandas().data
X = _clean(df[["age"]])
y = _make_y(df["censors"], df["survival"])
return X, y
def _load_democracy_duration():
df = ll_datasets.load_dd()
keep = ["start_year", "regime", "democracy"]
X = _clean(df[keep])
y = _make_y(df["observed"], df["duration"])
return X, y
# ── Registry ─────────────────────────────────────────────────────────────────
DATASET_LOADERS = {
"WHAS500": _load_whas500,
"GBSG2": _load_gbsg2,
"VeteransLungCancer": _load_veterans,
"FLChain": _load_flchain,
"AIDS": _load_aids,
"BreastCancerGenomic": _load_breast_cancer_genomic,
"Rossi": _load_rossi,
"KidneyTransplant": _load_kidney_transplant,
"NCTCGLung": _load_ncctg_lung,
"LymphNode": _load_lymph_node,
"Leukemia": _load_leukemia,
"Larynx": _load_larynx,
"Lymphoma": _load_lymphoma,
"StanfordHeart": _load_stanford_heart,
"DemocracyDuration": _load_democracy_duration,
}
DATASETS = list(DATASET_LOADERS.keys())
def load_dataset(name: str):
"""Returns (X_train, X_test, y_train, y_test) for dataset `name`."""
X, y = DATASET_LOADERS[name]()
X_tr, X_te, y_tr, y_te = _split(X, y)
return X_tr, X_te, y_tr, y_te
# Sanity check
print(f"{'Dataset':<25} {'train':>6} {'test':>6} {'features':>9} {'event_rate':>11}")
print("─" * 62)
for ds in DATASETS:
try:
Xtr, Xte, ytr, yte = load_dataset(ds)
er = np.mean([r[0] for r in ytr])
print(f"{ds:<25} {len(Xtr):>6} {len(Xte):>6} {Xtr.shape[1]:>9} {er:>11.2f}")
except Exception as e:
print(f"{ds:<25} ERROR: {e}")
print("\n✓ all datasets loaded")
"""## 3. Metric Helpers
We compute:
- **Harrell's C-index** (concordance index)
- **Brier Score** at the median event time (on test set)
- **Integrated Brier Score (IBS)** over the full range of test times
"""
from sksurv.metrics import (
concordance_index_censored,
brier_score as sksurv_brier_score,
integrated_brier_score as sksurv_ibs,
)
def _times_for_brier(y_train, y_test, n_points=20):
"""Evenly-spaced evaluation times within safe bounds."""
t_train_max = max(r[1] for r in y_train)
t_test_min = min(r[1] for r in y_test)
t_test_max = max(r[1] for r in y_test)
lo = t_test_min
hi = min(t_test_max * 0.95, t_train_max * 0.95)
if lo >= hi:
hi = lo * 1.5
return np.linspace(lo, hi, n_points)
def concordance_from_risk(risk_scores, y_test):
"""C-index given a 1-D array of risk scores (higher = more risk)."""
events = np.array([r[0] for r in y_test], dtype=bool)
times = np.array([r[1] for r in y_test])
c, _, _, _, _ = concordance_index_censored(events, times, risk_scores)
return c
def brier_ibs_from_surv_fns(surv_fns, y_train, y_test, times=None):
"""
Compute Brier score at median time and IBS.
surv_fns: list of step functions (scikit-survival API)
Returns (brier_at_median, ibs)
"""
if times is None:
times = _times_for_brier(y_train, y_test)
try:
preds = np.row_stack([
np.array([fn(t) for t in times]) for fn in surv_fns
]) # (n_subjects, n_times)
_, scores = sksurv_brier_score(y_train, y_test, preds, times)
brier_at_median = scores[len(times) // 2]
ibs = sksurv_ibs(y_train, y_test, preds, times)
return brier_at_median, ibs
except Exception:
return np.nan, np.nan
print("✓ metric helpers ok")
"""## 4. Collect all scikit-learn Regressors
We programmatically discover every sklearn regressor (excluding abstract, meta, or multioutput-only classes).
"""
from sklearn.utils import all_estimators
from sklearn.base import RegressorMixin
# Estimators known to require special setup / external deps — skip them
SKIP_REGRESSORS = {
"PLSRegression", # requires n_components <= n_features
"PLSCanonical",
"CCA",
"IsotonicRegression", # only 1-D input
"MultiOutputRegressor",
"RegressorChain",
"StackingRegressor",
"VotingRegressor",
"TransformedTargetRegressor",
"RadiusNeighborsRegressor", # can fail if no neighbors in radius
}
sklearn_regressors = []
for name, cls in all_estimators(type_filter="regressor"):
if name in SKIP_REGRESSORS:
continue
try:
cls() # check it can be instantiated with no args
sklearn_regressors.append((name, cls))
except Exception:
pass
print(f"Found {len(sklearn_regressors)} sklearn regressors to test")
print([n for n, _ in sklearn_regressors])
"""## 5. Benchmark `survivalist` with all sklearn Regressors
`SurvivalCustom` wraps any sklearn regressor for survival analysis.
All errors are caught — a failed model gets `NaN` metrics.
"""
from survivalist.custom import SurvivalCustom
def run_survivalist_model(regressor_cls, X_train, X_test, y_train, y_test):
"""
Defensive wrapper: fits SurvivalCustom(regressor_cls()) and returns metrics dict.
Returns NaNs on any failure.
"""
result = {
"c_index": np.nan,
"brier_median": np.nan,
"ibs": np.nan,
"fit_time_s": np.nan,
"error": None,
}
try:
model = SurvivalCustom(regr=regressor_cls(), random_state=123)
t0 = time.time()
model.fit(X_train, y_train)
result["fit_time_s"] = round(time.time() - t0, 3)
risk = model.predict(X_test)
result["c_index"] = round(concordance_from_risk(risk, y_test), 4)
surv_fns = model.predict_survival_function(X_test)
brier_med, ibs = brier_ibs_from_surv_fns(surv_fns, y_train, y_test)
result["brier_median"] = round(float(brier_med), 4) if not np.isnan(brier_med) else np.nan
result["ibs"] = round(float(ibs), 4) if not np.isnan(ibs) else np.nan
except Exception as e:
result["error"] = str(e)[:120]
return result
def benchmark_survivalist(dataset_name):
print(f"\n{'='*60}")
print(f" Dataset: {dataset_name}")
print(f"{'='*60}")
X_train, X_test, y_train, y_test = load_dataset(dataset_name)
rows = []
for reg_name, reg_cls in tqdm(sklearn_regressors, desc=dataset_name):
metrics = run_survivalist_model(reg_cls, X_train, X_test, y_train, y_test)
rows.append({
"dataset": dataset_name,
"package": "survivalist",
"model": f"SurvivalCustom({reg_name})",
"c_index": metrics["c_index"],
"brier_median": metrics["brier_median"],
"ibs": metrics["ibs"],
"fit_time_s": metrics["fit_time_s"],
"error": metrics["error"],
})
status = "✓" if metrics["error"] is None else "✗"
print(
f" {status} {reg_name:45s} "
f"C-index={metrics['c_index']} IBS={metrics['ibs']} "
f"t={metrics['fit_time_s']}s"
)
return rows
# ── Run across all 15 datasets ────────────────────────────────────────────────
all_rows = []
for ds in DATASETS:
all_rows.extend(benchmark_survivalist(ds))
survivalist_df = pd.DataFrame(all_rows)
print(f"\nTotal experiments: {len(survivalist_df)}")
print(f"Successful: {survivalist_df['error'].isna().sum()}")
print(f"Failed: {survivalist_df['error'].notna().sum()}")
"""## 6. Baseline Models: `scikit-survival` + `lifelines`"""
# ── scikit-survival baselines ─────────────────────────────────────────────────
from sksurv.linear_model import CoxPHSurvivalAnalysis
from sksurv.ensemble import RandomSurvivalForest, GradientBoostingSurvivalAnalysis
# ── lifelines baselines ───────────────────────────────────────────────────────
from lifelines import CoxPHFitter
def run_sksurv_model(model, X_train, X_test, y_train, y_test):
result = {"c_index": np.nan, "brier_median": np.nan, "ibs": np.nan,
"fit_time_s": np.nan, "error": None}
try:
t0 = time.time()
model.fit(X_train, y_train)
result["fit_time_s"] = round(time.time() - t0, 3)
risk = model.predict(X_test)
result["c_index"] = round(concordance_from_risk(risk, y_test), 4)
times = _times_for_brier(y_train, y_test)
try:
surv_fns = model.predict_survival_function(X_test)
bm, ibs = brier_ibs_from_surv_fns(surv_fns, y_train, y_test, times)
result["brier_median"] = round(float(bm), 4) if not np.isnan(bm) else np.nan
result["ibs"] = round(float(ibs), 4) if not np.isnan(ibs) else np.nan
except Exception:
pass
except Exception as e:
result["error"] = str(e)[:120]
return result
def run_lifelines_cox(X_train, X_test, y_train, y_test):
result = {"c_index": np.nan, "brier_median": np.nan, "ibs": np.nan,
"fit_time_s": np.nan, "error": None}
try:
df_train = X_train.copy()
df_train["duration"] = [r[1] for r in y_train]
df_train["event"] = [int(r[0]) for r in y_train]
df_test = X_test.copy()
df_test["duration"] = [r[1] for r in y_test]
df_test["event"] = [int(r[0]) for r in y_test]
cph = CoxPHFitter(penalizer=0.1)
t0 = time.time()
cph.fit(df_train, duration_col="duration", event_col="event")
result["fit_time_s"] = round(time.time() - t0, 3)
result["c_index"] = round(cph.score(df_test, scoring_method="concordance_index"), 4)
except Exception as e:
result["error"] = str(e)[:120]
return result
def benchmark_baselines(dataset_name):
print(f"\n── Baselines: {dataset_name} ──")
X_train, X_test, y_train, y_test = load_dataset(dataset_name)
baselines = [
("sksurv", "CoxPHSurvivalAnalysis",
lambda: run_sksurv_model(CoxPHSurvivalAnalysis(alpha=0.1), X_train, X_test, y_train, y_test)),
("sksurv", "RandomSurvivalForest",
lambda: run_sksurv_model(RandomSurvivalForest(n_estimators=100, random_state=RANDOM_STATE),
X_train, X_test, y_train, y_test)),
("sksurv", "GradientBoostingSurvivalAnalysis",
lambda: run_sksurv_model(GradientBoostingSurvivalAnalysis(random_state=RANDOM_STATE),
X_train, X_test, y_train, y_test)),
("lifelines", "CoxPHFitter",
lambda: run_lifelines_cox(X_train, X_test, y_train, y_test)),
]
rows = []
for pkg, mname, fn in baselines:
metrics = fn()
status = "✓" if metrics["error"] is None else "✗"
print(f" {status} {pkg}/{mname:40s} C-index={metrics['c_index']} "
f"IBS={metrics['ibs']} t={metrics['fit_time_s']}s")
rows.append({
"dataset": dataset_name,
"package": pkg,
"model": mname,
"c_index": metrics["c_index"],
"brier_median": metrics["brier_median"],
"ibs": metrics["ibs"],
"fit_time_s": metrics["fit_time_s"],
"error": metrics["error"],
})
return rows
baseline_rows = []
for ds in DATASETS:
baseline_rows.extend(benchmark_baselines(ds))
baseline_df = pd.DataFrame(baseline_rows)
print("\n✓ baselines done")
"""## 7. Combine Results & Save"""
results_df = pd.concat([survivalist_df, baseline_df], ignore_index=True)
results_df["success"] = results_df["error"].isna()
results_df.to_csv("survival_benchmark_results.csv", index=False)
print(f"Saved {len(results_df)} rows to survival_benchmark_results.csv")
results_df.head(10)
"""## 8. Results Tables"""
def pretty_table(dataset_name):
df = results_df[results_df["dataset"] == dataset_name].copy()
df = df[df["success"]].sort_values("c_index", ascending=False)
display_cols = ["package", "model", "c_index", "brier_median", "ibs", "fit_time_s"]
print(f"\n{'═'*100}")
print(f" {dataset_name} — successful models only (sorted by C-index ↓)")
print(f"{'═'*100}")
print(df[display_cols].to_string(index=False))
return df
for ds in DATASETS:
pretty_table(ds)
"""## 9. Aggregate Statistics Across All Datasets
We compute per-dataset mean/median C-index for `survivalist` vs baselines, and rank models by their aggregate performance.
"""
# ── Per-dataset summary ───────────────────────────────────────────────────────
summary_rows = []
for ds in DATASETS:
sub = results_df[(results_df["dataset"] == ds) & results_df["success"]]
surv = sub[sub["package"] == "survivalist"]["c_index"].dropna()
base = sub[sub["package"] != "survivalist"]
best_base_c = base["c_index"].max()
best_surv_c = surv.max()
median_surv_c = surv.median()
summary_rows.append({
"dataset": ds,
"n_survivalist_ok": len(surv),
"survivalist_mean_C": round(surv.mean(), 4),
"survivalist_median_C":round(surv.median(), 4),
"survivalist_max_C": round(surv.max(), 4),
"best_baseline_C": round(best_base_c, 4) if not np.isnan(best_base_c) else np.nan,
"survivalist_beats_best_baseline": bool(best_surv_c > best_base_c),
})
summary_df = pd.DataFrame(summary_rows)
print("\n── Cross-dataset summary ──")
print(summary_df.to_string(index=False))
wins = summary_df["survivalist_beats_best_baseline"].sum()
print(f"\nsurvivalist best-model beats best baseline on {wins}/{len(DATASETS)} datasets")
"""## 10. Failure Summary"""
failed = results_df[~results_df["success"] & results_df["package"].eq("survivalist")]
if len(failed):
print(f"Survivalist failures: {len(failed)}")
for _, row in failed.iterrows():
print(f" [{row['dataset']}] {row['model']}: {row['error']}")
else:
print("No survivalist failures — all models ran successfully!")
# Failure rate by dataset
print("\n── Failure rate by dataset ──")
for ds in DATASETS:
sub = results_df[(results_df["dataset"] == ds) & (results_df["package"] == "survivalist")]
n_fail = (~sub["success"]).sum()
print(f" {ds:<25} {n_fail:>3} / {len(sub)} failed")
"""## 11. Visualisations
### 11.1 C-index Distribution — survivalist vs baselines (all 15 datasets)
"""
n_ds = len(DATASETS)
ncols = 3
nrows = (n_ds + ncols - 1) // ncols
fig, axes = plt.subplots(nrows, ncols, figsize=(7 * ncols, 5 * nrows))
axes = axes.flatten()
for ax, ds in zip(axes, DATASETS):
sub = results_df[(results_df["dataset"] == ds) & results_df["success"]].copy()
surv = sub[sub["package"] == "survivalist"]["c_index"].dropna()
base = sub[sub["package"] != "survivalist"][["model", "c_index"]].dropna()
if len(surv) > 1:
ax.violinplot([surv], positions=[0], showmedians=True)
ax.scatter(
np.zeros(len(surv)) + np.random.normal(0, 0.02, len(surv)),
surv, alpha=0.35, s=10, color="steelblue",
)
colors = plt.cm.tab10(np.linspace(0, 1, max(1, len(base))))
for i, (_, r) in enumerate(base.iterrows()):
ax.axhline(r["c_index"], linestyle="--", color=colors[i],
linewidth=1.5, label=r["model"])
ax.set_title(ds, fontsize=10)
ax.set_xticks([0])
ax.set_xticklabels(["survivalist"])
ax.set_ylabel("C-index")
ax.legend(fontsize=6, loc="lower right")
ax.set_ylim(0.3, 1.02)
# Hide unused axes
for ax in axes[n_ds:]:
ax.set_visible(False)
plt.suptitle("C-index: survivalist (all sklearn wrappers) vs established baselines — 15 datasets",
fontsize=13)
plt.tight_layout()
plt.savefig("c_index_violin_all_datasets.png", dpi=150, bbox_inches="tight")
plt.show()
"""### 11.2 Integrated Brier Score Distribution (all 15 datasets)"""
fig, axes = plt.subplots(nrows, ncols, figsize=(7 * ncols, 5 * nrows))
axes = axes.flatten()
for ax, ds in zip(axes, DATASETS):
sub = results_df[(results_df["dataset"] == ds) & results_df["success"]].copy()
surv = sub[sub["package"] == "survivalist"]["ibs"].dropna()
base = sub[sub["package"] != "survivalist"][["model", "ibs"]].dropna()
if len(surv) > 1:
ax.violinplot([surv], positions=[0], showmedians=True)
ax.scatter(
np.zeros(len(surv)) + np.random.normal(0, 0.02, len(surv)),
surv, alpha=0.35, s=10, color="darkorange",
)
colors = plt.cm.tab10(np.linspace(0, 1, max(1, len(base))))
for i, (_, r) in enumerate(base.iterrows()):
ax.axhline(r["ibs"], linestyle="--", color=colors[i],
linewidth=1.5, label=r["model"])
ax.set_title(ds, fontsize=10)
ax.set_xticks([0])
ax.set_xticklabels(["survivalist"])
ax.set_ylabel("IBS (lower=better)")
ax.legend(fontsize=6)
for ax in axes[n_ds:]:
ax.set_visible(False)
plt.suptitle("Integrated Brier Score: survivalist vs baselines — 15 datasets", fontsize=13)
plt.tight_layout()
plt.savefig("ibs_violin_all_datasets.png", dpi=150, bbox_inches="tight")
plt.show()
"""### 11.3 Top-10 survivalist models per dataset (C-index)"""
fig, axes = plt.subplots(nrows, ncols, figsize=(8 * ncols, 5 * nrows))
axes = axes.flatten()
for ax, ds in zip(axes, DATASETS):
sub = results_df[
(results_df["dataset"] == ds)
& results_df["success"]
& (results_df["package"] == "survivalist")
].copy()
top10 = sub.nlargest(10, "c_index")
labels = (top10["model"]
.str.replace("SurvivalCustom(", "", regex=False)
.str.replace(")", "", regex=False))
bars = ax.barh(range(len(top10)), top10["c_index"].values,
color="steelblue", edgecolor="white")
ax.set_yticks(range(len(top10)))
ax.set_yticklabels(labels, fontsize=7)
ax.invert_yaxis()
ax.set_xlabel("C-index")
ax.set_title(f"{ds} — Top 10 by C-index", fontsize=10)
ax.set_xlim(0.4, 1.02)
for bar, val in zip(bars, top10["c_index"].values):
ax.text(val + 0.002, bar.get_y() + bar.get_height() / 2,
f"{val:.3f}", va="center", fontsize=7)
for ax in axes[n_ds:]:
ax.set_visible(False)
plt.suptitle("Top 10 survivalist/sklearn models per dataset (C-index)", fontsize=13)
plt.tight_layout()
plt.savefig("top10_cindex_all_datasets.png", dpi=150, bbox_inches="tight")
plt.show()
"""### 11.4 Heatmap: Median C-index by Regressor × Dataset"""
# Pivot: rows = regressor, cols = dataset, values = c_index
surv_only = results_df[
results_df["success"] & (results_df["package"] == "survivalist")
].copy()
surv_only["regressor"] = (surv_only["model"]
.str.replace("SurvivalCustom(", "", regex=False)
.str.replace(")", "", regex=False))
pivot = surv_only.pivot_table(
index="regressor", columns="dataset", values="c_index", aggfunc="mean"
)
# Keep only regressors that succeeded on at least half the datasets
pivot = pivot[pivot.notna().sum(axis=1) >= len(DATASETS) // 2]
fig, ax = plt.subplots(figsize=(len(DATASETS) * 1.0, max(8, len(pivot) * 0.35)))
sns.heatmap(
pivot, ax=ax, cmap="RdYlGn", center=0.65,
linewidths=0.3, linecolor="white",
annot=True, fmt=".2f", annot_kws={"size": 6},
vmin=0.40, vmax=0.95,
)
ax.set_title("Median C-index by Regressor × Dataset\n(survivalist wrappers)", fontsize=12)
ax.set_xlabel("")
ax.set_ylabel("")
ax.tick_params(axis="x", rotation=40, labelsize=8)
ax.tick_params(axis="y", labelsize=7)
plt.tight_layout()
plt.savefig("heatmap_regressor_dataset.png", dpi=150, bbox_inches="tight")
plt.show()
"""### 11.5 Aggregate Ranking: Mean C-index Across All Datasets"""
mean_c = (surv_only.groupby("regressor")["c_index"]
.mean()
.sort_values(ascending=False)
.dropna())
fig, ax = plt.subplots(figsize=(9, max(6, len(mean_c) * 0.3)))
bars = ax.barh(range(len(mean_c)), mean_c.values, color="steelblue", edgecolor="white")
ax.set_yticks(range(len(mean_c)))
ax.set_yticklabels(mean_c.index, fontsize=8)
ax.invert_yaxis()
ax.set_xlabel("Mean C-index across all datasets")
ax.set_title("Aggregate ranking: survivalist/sklearn regressors — all 15 datasets", fontsize=11)
ax.set_xlim(0.40, mean_c.max() + 0.04)
for bar, val in zip(bars, mean_c.values):
ax.text(val + 0.001, bar.get_y() + bar.get_height() / 2,
f"{val:.3f}", va="center", fontsize=7)
plt.tight_layout()
plt.savefig("aggregate_ranking.png", dpi=150, bbox_inches="tight")
plt.show()
"""### 11.6 Cross-dataset Summary: survivalist vs Best Baseline"""
fig, ax = plt.subplots(figsize=(10, 5))
x = np.arange(len(DATASETS))
width = 0.35
surv_c = summary_df["survivalist_max_C"].values
base_c = summary_df["best_baseline_C"].values
ax.bar(x - width/2, surv_c, width, label="survivalist (best)", color="steelblue", alpha=0.85)
ax.bar(x + width/2, base_c, width, label="Best baseline", color="salmon", alpha=0.85)
ax.set_xticks(x)
ax.set_xticklabels(DATASETS, rotation=35, ha="right", fontsize=8)
ax.set_ylabel("C-index")
ax.set_title("Best survivalist model vs Best baseline — all 15 datasets")
ax.legend()
ax.axhline(0.5, color="grey", linestyle=":", linewidth=0.8)
ax.set_ylim(0.3, 1.05)
plt.tight_layout()
plt.savefig("survivalist_vs_baseline_summary.png", dpi=150, bbox_inches="tight")
plt.show()
print("\n── Win/loss summary ──")
print(summary_df[["dataset", "survivalist_max_C", "best_baseline_C",
"survivalist_beats_best_baseline"]].to_string(index=False))
… ✓ base imports ok Dataset train test features event_rate ────────────────────────────────────────────────────────────── WHAS500 375 125 22 0.43 GBSG2 514 172 12 0.44 VeteransLungCancer 102 35 11 0.92 FLChain 5905 1969 43 0.27 AIDS 863 288 27 0.08 BreastCancerGenomic 148 50 84 0.22 Rossi 324 108 7 0.25 KidneyTransplant 647 216 4 0.16 NCTCGLung 125 42 8 0.31 LymphNode 514 172 8 0.23 Leukemia 31 11 3 0.71 Larynx 67 23 4 0.55 Lymphoma 60 20 1 0.65 StanfordHeart 51 18 1 0.59 DemocracyDuration 1356 452 9 0.81
✓ all datasets loaded
✓ metric helpers ok
Found 45 sklearn regressors to test
['ARDRegression', 'AdaBoostRegressor', 'BaggingRegressor', 'BayesianRidge', 'DecisionTreeRegressor', 'DummyRegressor', 'ElasticNet', 'ElasticNetCV', 'ExtraTreeRegressor', 'ExtraTreesRegressor', 'GammaRegressor', 'GaussianProcessRegressor', 'GradientBoostingRegressor', 'HistGradientBoostingRegressor', 'HuberRegressor', 'KNeighborsRegressor', 'KernelRidge', 'Lars', 'LarsCV', 'Lasso', 'LassoCV', 'LassoLars', 'LassoLarsCV', 'LassoLarsIC', 'LinearRegression', 'LinearSVR', 'MLPRegressor', 'MultiTaskElasticNet', 'MultiTaskElasticNetCV', 'MultiTaskLasso', 'MultiTaskLassoCV', 'NuSVR', 'OrthogonalMatchingPursuit', 'OrthogonalMatchingPursuitCV', 'PassiveAggressiveRegressor', 'PoissonRegressor', 'QuantileRegressor', 'RANSACRegressor', 'RandomForestRegressor', 'Ridge', 'RidgeCV', 'SGDRegressor', 'SVR', 'TheilSenRegressor', 'TweedieRegressor']
============================================================
Dataset: WHAS500
============================================================
WHAS500: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.717 IBS=0.3011 t=0.148s
✓ AdaBoostRegressor C-index=0.6819 IBS=0.3025 t=1.809s
✓ BaggingRegressor C-index=0.6725 IBS=0.3496 t=1.583s
✓ BayesianRidge C-index=0.6873 IBS=0.3218 t=0.088s
✓ DecisionTreeRegressor C-index=0.6251 IBS=0.3495 t=0.128s
✓ DummyRegressor C-index=0.5 IBS=0.2338 t=0.018s
✓ ElasticNet C-index=0.6787 IBS=0.303 t=0.036s
✓ ElasticNetCV C-index=0.7163 IBS=0.3072 t=1.359s
✓ ExtraTreeRegressor C-index=0.6242 IBS=0.363 t=0.069s
✓ ExtraTreesRegressor C-index=0.7462 IBS=0.3434 t=6.299s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.5005 IBS=0.3858 t=1.071s
✓ GradientBoostingRegressor C-index=0.6986 IBS=0.338 t=4.199s
✓ HistGradientBoostingRegressor C-index=0.6818 IBS=0.3788 t=2.73s
✓ HuberRegressor C-index=0.7193 IBS=0.339 t=0.952s
✓ KNeighborsRegressor C-index=0.6774 IBS=0.3095 t=0.059s
✓ KernelRidge C-index=0.7208 IBS=0.3152 t=0.433s
✓ Lars C-index=0.7199 IBS=0.3143 t=0.114s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.6795 IBS=0.2815 t=0.036s
✓ LassoCV C-index=0.7168 IBS=0.3072 t=1.225s
✓ LassoLars C-index=0.6795 IBS=0.2815 t=0.05s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.7174 IBS=0.3048 t=0.114s
✓ LinearRegression C-index=0.7189 IBS=0.316 t=0.044s
✓ LinearSVR C-index=0.7246 IBS=0.3753 t=0.587s
✓ MLPRegressor C-index=0.6431 IBS=0.4129 t=7.804s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.6846 IBS=0.281 t=0.287s
✓ OrthogonalMatchingPursuit C-index=0.6826 IBS=0.277 t=0.029s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.7072 IBS=0.4363 t=0.046s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.681 IBS=0.3223 t=0.65s
✓ RANSACRegressor C-index=0.6876 IBS=0.39 t=2.657s
✓ RandomForestRegressor C-index=0.714 IBS=0.3285 t=8.702s
✓ Ridge C-index=0.7191 IBS=0.3145 t=0.043s
✓ RidgeCV C-index=0.7187 IBS=0.3117 t=0.095s
✓ SGDRegressor C-index=0.4395 IBS=0.6711 t=0.131s
✓ SVR C-index=0.6806 IBS=0.3555 t=0.406s
✓ TheilSenRegressor C-index=0.7174 IBS=0.3144 t=26.558s
✓ TweedieRegressor C-index=0.6962 IBS=0.3129 t=0.477s
============================================================
Dataset: GBSG2
============================================================
GBSG2: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.7098 IBS=0.3462 t=0.076s
✓ AdaBoostRegressor C-index=0.6447 IBS=0.2121 t=0.75s
✓ BaggingRegressor C-index=0.6114 IBS=0.2722 t=0.602s
✓ BayesianRidge C-index=0.6502 IBS=0.3168 t=0.038s
✓ DecisionTreeRegressor C-index=0.584 IBS=0.277 t=0.067s
✓ DummyRegressor C-index=0.5 IBS=0.1916 t=0.019s
✓ ElasticNet C-index=0.6565 IBS=0.233 t=0.028s
✓ ElasticNetCV C-index=0.6486 IBS=0.3082 t=0.712s
✓ ExtraTreeRegressor C-index=0.5961 IBS=0.2942 t=0.045s
✓ ExtraTreesRegressor C-index=0.6221 IBS=0.279 t=4.611s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.5565 IBS=0.3011 t=1.237s
✓ GradientBoostingRegressor C-index=0.6228 IBS=0.257 t=1.528s
✓ HistGradientBoostingRegressor C-index=0.5952 IBS=0.2776 t=1.806s
✓ HuberRegressor C-index=0.702 IBS=0.3873 t=0.496s
✓ KNeighborsRegressor C-index=0.5529 IBS=0.2697 t=0.078s
✓ KernelRidge C-index=0.684 IBS=0.3387 t=0.46s
✓ Lars C-index=0.6833 IBS=0.3384 t=0.046s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.615 IBS=0.1912 t=0.027s
✓ LassoCV C-index=0.6475 IBS=0.3064 t=0.68s
✓ LassoLars C-index=0.615 IBS=0.1912 t=0.031s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.6833 IBS=0.3384 t=0.071s
✓ LinearRegression C-index=0.6833 IBS=0.3384 t=0.029s
✓ LinearSVR C-index=0.6137 IBS=0.4824 t=0.456s
✓ MLPRegressor C-index=0.5739 IBS=0.4639 t=5.809s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.6062 IBS=0.2051 t=0.35s
✓ OrthogonalMatchingPursuit C-index=0.6071 IBS=0.1899 t=0.025s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.6191 IBS=0.3391 t=0.027s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.6071 IBS=0.1899 t=0.437s
✓ RANSACRegressor C-index=0.6523 IBS=0.5078 t=1.412s
✓ RandomForestRegressor C-index=0.6129 IBS=0.2625 t=4.755s
✓ Ridge C-index=0.6833 IBS=0.3386 t=0.05s
✓ RidgeCV C-index=0.6832 IBS=0.3399 t=0.052s
✓ SGDRegressor C-index=0.5508 IBS=0.3983 t=0.063s
✓ SVR C-index=0.6044 IBS=0.2354 t=0.761s
✓ TheilSenRegressor C-index=0.6742 IBS=0.4324 t=7.961s
✓ TweedieRegressor C-index=0.6658 IBS=0.3398 t=0.235s
============================================================
Dataset: VeteransLungCancer
============================================================
VeteransLungCancer: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.627 IBS=nan t=0.125s
✓ AdaBoostRegressor C-index=0.6243 IBS=nan t=1.045s
✓ BaggingRegressor C-index=0.6052 IBS=nan t=0.461s
✓ BayesianRidge C-index=0.7026 IBS=nan t=0.084s
✓ DecisionTreeRegressor C-index=0.6426 IBS=nan t=0.029s
✓ DummyRegressor C-index=0.5 IBS=nan t=0.013s
✓ ElasticNet C-index=0.7078 IBS=nan t=0.036s
✓ ElasticNetCV C-index=0.6557 IBS=nan t=0.896s
✓ ExtraTreeRegressor C-index=0.607 IBS=nan t=0.034s
✓ ExtraTreesRegressor C-index=0.6209 IBS=nan t=1.917s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.5087 IBS=nan t=0.046s
✓ GradientBoostingRegressor C-index=0.6035 IBS=nan t=0.761s
✓ HistGradientBoostingRegressor C-index=0.6487 IBS=nan t=0.531s
✓ HuberRegressor C-index=0.6574 IBS=nan t=0.383s
✓ KNeighborsRegressor C-index=0.6443 IBS=nan t=0.037s
✓ KernelRidge C-index=0.633 IBS=nan t=0.018s
✓ Lars C-index=0.6383 IBS=nan t=0.035s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.7078 IBS=nan t=0.014s
✓ LassoCV C-index=0.6504 IBS=nan t=0.646s
✓ LassoLars C-index=0.7078 IBS=nan t=0.017s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.6452 IBS=nan t=0.045s
✓ LinearRegression C-index=0.6383 IBS=nan t=0.015s
✓ LinearSVR C-index=0.6313 IBS=nan t=0.069s
✓ MLPRegressor C-index=0.6539 IBS=nan t=0.619s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.7061 IBS=nan t=0.038s
✓ OrthogonalMatchingPursuit C-index=0.7078 IBS=nan t=0.011s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.6365 IBS=nan t=0.02s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.7078 IBS=nan t=0.1s
✓ RANSACRegressor C-index=0.6 IBS=nan t=1.426s
✓ RandomForestRegressor C-index=0.6504 IBS=nan t=1.8s
✓ Ridge C-index=0.6365 IBS=nan t=0.016s
✓ RidgeCV C-index=0.6713 IBS=nan t=0.027s
✓ SGDRegressor C-index=0.5617 IBS=nan t=0.019s
✓ SVR C-index=0.7078 IBS=nan t=0.025s
✓ TheilSenRegressor C-index=0.647 IBS=nan t=7.139s
✓ TweedieRegressor C-index=0.7009 IBS=nan t=0.198s
============================================================
Dataset: FLChain
============================================================
FLChain: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.9223 IBS=0.1201 t=1.259s
✓ AdaBoostRegressor C-index=0.8854 IBS=0.1283 t=5.704s
✓ BaggingRegressor C-index=0.9304 IBS=0.1176 t=24.055s
✓ BayesianRidge C-index=0.9299 IBS=0.1205 t=1.498s
✓ DecisionTreeRegressor C-index=0.9229 IBS=0.119 t=3.875s
✓ DummyRegressor C-index=0.5 IBS=0.1305 t=0.662s
✓ ElasticNet C-index=0.7722 IBS=0.1413 t=1.199s
✓ ElasticNetCV C-index=0.9267 IBS=0.1766 t=5.788s
✓ ExtraTreeRegressor C-index=0.9229 IBS=0.119 t=2.492s
✓ ExtraTreesRegressor C-index=0.9322 IBS=0.1185 t=184.103s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.8761 IBS=0.1352 t=556.617s
✓ GradientBoostingRegressor C-index=0.9279 IBS=0.1138 t=42.219s
✓ HistGradientBoostingRegressor C-index=0.9274 IBS=0.1414 t=26.807s
✓ HuberRegressor C-index=0.9193 IBS=0.2541 t=17.314s
✓ KNeighborsRegressor C-index=0.8074 IBS=0.1462 t=13.201s
✓ KernelRidge C-index=0.9312 IBS=0.1276 t=176.373s
✓ Lars C-index=0.9288 IBS=0.1424 t=1.288s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.7722 IBS=0.1329 t=0.851s
✓ LassoCV C-index=0.9274 IBS=0.1663 t=8.204s
✓ LassoLars C-index=0.7722 IBS=0.1329 t=1.001s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.9299 IBS=0.1205 t=4.031s
✓ LinearRegression C-index=0.9299 IBS=0.1205 t=3.134s
✓ LinearSVR C-index=0.9289 IBS=0.1247 t=19.291s
✓ MLPRegressor C-index=0.9304 IBS=0.1367 t=62.198s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.861 IBS=0.2591 t=160.384s
✓ OrthogonalMatchingPursuit C-index=0.8876 IBS=0.2241 t=0.898s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.9283 IBS=0.1424 t=5.929s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.7722 IBS=0.13 t=75.045s
✓ RANSACRegressor C-index=0.8328 IBS=0.1574 t=12.349s
✓ RandomForestRegressor C-index=0.9309 IBS=0.1158 t=225.531s
✓ Ridge C-index=0.9312 IBS=0.128 t=1.107s
✓ RidgeCV C-index=0.9301 IBS=0.1206 t=2.316s
✓ SGDRegressor C-index=0.5146 IBS=0.8268 t=8.477s
✓ SVR C-index=0.8599 IBS=0.263 t=149.16s
✓ TheilSenRegressor C-index=0.9225 IBS=0.135 t=214.08s
✓ TweedieRegressor C-index=0.8094 IBS=0.176 t=1.138s
============================================================
Dataset: AIDS
============================================================
AIDS: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.7366 IBS=0.0725 t=0.247s
✓ AdaBoostRegressor C-index=0.75 IBS=0.0784 t=0.783s
✓ BaggingRegressor C-index=0.6309 IBS=0.08 t=1.823s
✓ BayesianRidge C-index=0.76 IBS=0.0728 t=0.158s
✓ DecisionTreeRegressor C-index=0.5148 IBS=0.0798 t=0.242s
✓ DummyRegressor C-index=0.5 IBS=0.0739 t=0.04s
✓ ElasticNet C-index=0.763 IBS=0.0733 t=0.06s
✓ ElasticNetCV C-index=0.761 IBS=0.0727 t=2.761s
✓ ExtraTreeRegressor C-index=0.4762 IBS=0.089 t=0.213s
✓ ExtraTreesRegressor C-index=0.6429 IBS=0.0795 t=14.052s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.5034 IBS=0.0797 t=12.94s
✓ GradientBoostingRegressor C-index=0.6962 IBS=0.0758 t=4.841s
✓ HistGradientBoostingRegressor C-index=0.6587 IBS=0.0756 t=8.208s
✓ HuberRegressor C-index=0.572 IBS=0.074 t=1.691s
✓ KNeighborsRegressor C-index=0.6527 IBS=0.0726 t=0.212s
✓ KernelRidge C-index=0.7424 IBS=0.0723 t=2.222s
✓ Lars C-index=0.736 IBS=0.0724 t=0.228s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.763 IBS=0.0734 t=0.053s
✓ LassoCV C-index=0.761 IBS=0.0727 t=2.251s
✓ LassoLars C-index=0.763 IBS=0.0734 t=0.08s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.7536 IBS=0.0725 t=1.074s
✓ LinearRegression C-index=0.7415 IBS=0.0723 t=0.103s
✓ LinearSVR C-index=0.6687 IBS=0.0726 t=2.106s
✓ MLPRegressor C-index=0.7139 IBS=0.0736 t=27.167s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.5596 IBS=0.074 t=2.886s
✓ OrthogonalMatchingPursuit C-index=0.7306 IBS=0.0731 t=0.064s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.7147 IBS=0.0724 t=0.081s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.5 IBS=0.0741 t=1.84s
✓ RANSACRegressor C-index=0.526 IBS=0.074 t=3.449s
✓ RandomForestRegressor C-index=0.6726 IBS=0.0752 t=17.432s
✓ Ridge C-index=0.7422 IBS=0.0723 t=0.08s
✓ RidgeCV C-index=0.7461 IBS=0.0724 t=0.128s
✓ SGDRegressor C-index=0.7398 IBS=0.086 t=0.28s
✓ SVR C-index=0.6709 IBS=0.0738 t=0.978s
✓ TheilSenRegressor C-index=0.7399 IBS=0.0732 t=42.094s
✓ TweedieRegressor C-index=0.7518 IBS=0.073 t=0.478s
============================================================
Dataset: BreastCancerGenomic
============================================================
BreastCancerGenomic: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.6455 IBS=0.2353 t=7.201s
✓ AdaBoostRegressor C-index=0.6476 IBS=0.2305 t=18.053s
✓ BaggingRegressor C-index=0.4579 IBS=0.286 t=7.901s
✓ BayesianRidge C-index=0.6359 IBS=0.2271 t=1.312s
✓ DecisionTreeRegressor C-index=0.5579 IBS=0.3154 t=0.73s
✓ DummyRegressor C-index=0.5 IBS=0.2379 t=0.042s
✓ ElasticNet C-index=0.5 IBS=0.2379 t=0.092s
✓ ElasticNetCV C-index=0.5903 IBS=0.2308 t=26.208s
✓ ExtraTreeRegressor C-index=0.4703 IBS=0.2868 t=0.338s
✓ ExtraTreesRegressor C-index=0.6497 IBS=0.263 t=31.125s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.5 IBS=0.2742 t=0.876s
✓ GradientBoostingRegressor C-index=0.5393 IBS=0.2844 t=33.544s
✓ HistGradientBoostingRegressor C-index=0.6538 IBS=0.2583 t=12.298s
✓ HuberRegressor C-index=0.6221 IBS=0.2731 t=3.563s
✓ KNeighborsRegressor C-index=0.5317 IBS=0.2407 t=0.131s
✓ KernelRidge C-index=0.6166 IBS=0.3 t=0.748s
✓ Lars C-index=0.5959 IBS=0.3238 t=1.879s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.5 IBS=0.2379 t=0.118s
✓ LassoCV C-index=0.589 IBS=0.2308 t=27.687s
✓ LassoLars C-index=0.5 IBS=0.2379 t=0.445s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.5834 IBS=0.2309 t=4.653s
✓ LinearRegression C-index=0.6014 IBS=0.3157 t=0.364s
✓ LinearSVR C-index=0.6331 IBS=0.267 t=3.115s
✓ MLPRegressor C-index=0.5862 IBS=0.2658 t=16.889s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.6193 IBS=0.2366 t=0.337s
✓ OrthogonalMatchingPursuit C-index=0.5214 IBS=0.2509 t=0.234s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.6303 IBS=0.2296 t=0.212s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.5 IBS=0.2449 t=2.956s
✓ RANSACRegressor C-index=0.3821 IBS=0.7496 t=22.216s
✓ RandomForestRegressor C-index=0.5241 IBS=0.2503 t=67.673s
✓ Ridge C-index=0.6152 IBS=0.302 t=0.211s
✓ RidgeCV C-index=0.6317 IBS=0.2596 t=0.443s
✓ SGDRegressor C-index=0.4 IBS=0.9192 t=0.286s
✓ SVR C-index=0.6317 IBS=0.2351 t=0.281s
✓ TheilSenRegressor C-index=0.6428 IBS=0.3167 t=1618.689s
✓ TweedieRegressor C-index=0.6441 IBS=0.2274 t=1.736s
============================================================
Dataset: Rossi
============================================================
Rossi: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.5559 IBS=0.1108 t=0.05s
✓ AdaBoostRegressor C-index=0.5979 IBS=0.1162 t=0.142s
✓ BaggingRegressor C-index=0.5996 IBS=0.1278 t=0.233s
✓ BayesianRidge C-index=0.675 IBS=0.1094 t=0.035s
✓ DecisionTreeRegressor C-index=0.5699 IBS=0.1441 t=0.02s
✓ DummyRegressor C-index=0.5 IBS=0.1117 t=0.007s
✓ ElasticNet C-index=0.5 IBS=0.1117 t=0.018s
✓ ElasticNetCV C-index=0.624 IBS=0.11 t=0.438s
✓ ExtraTreeRegressor C-index=0.5516 IBS=0.1506 t=0.019s
✓ ExtraTreesRegressor C-index=0.5689 IBS=0.1368 t=1.135s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.5978 IBS=0.1317 t=0.128s
✓ GradientBoostingRegressor C-index=0.6186 IBS=0.1346 t=0.587s
✓ HistGradientBoostingRegressor C-index=0.5966 IBS=0.1154 t=0.726s
✓ HuberRegressor C-index=0.5836 IBS=0.1137 t=0.403s
✓ KNeighborsRegressor C-index=0.6121 IBS=0.1149 t=0.042s
✓ KernelRidge C-index=0.5689 IBS=0.1115 t=0.066s
✓ Lars C-index=0.5686 IBS=0.1115 t=0.031s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.5 IBS=0.1117 t=0.021s
✓ LassoCV C-index=0.6173 IBS=0.11 t=0.615s
✓ LassoLars C-index=0.5 IBS=0.1117 t=0.027s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.6036 IBS=0.1102 t=0.046s
✓ LinearRegression C-index=0.5686 IBS=0.1115 t=0.021s
✓ LinearSVR C-index=0.5619 IBS=0.1088 t=0.186s
✓ MLPRegressor C-index=0.5799 IBS=0.1365 t=1.641s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.6106 IBS=0.1114 t=0.115s
✓ OrthogonalMatchingPursuit C-index=0.6603 IBS=0.1105 t=0.022s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.5873 IBS=0.1101 t=0.023s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.5 IBS=0.1143 t=0.132s
✓ RANSACRegressor C-index=0.5 IBS=0.1143 t=0.571s
✓ RandomForestRegressor C-index=0.6054 IBS=0.1237 t=1.335s
✓ Ridge C-index=0.5706 IBS=0.1115 t=0.014s
✓ RidgeCV C-index=0.5792 IBS=0.1111 t=0.022s
✓ SGDRegressor C-index=0.5379 IBS=0.2751 t=0.085s
✓ SVR C-index=0.6143 IBS=0.1124 t=0.114s
✓ TheilSenRegressor C-index=0.6006 IBS=0.1095 t=3.562s
✓ TweedieRegressor C-index=0.661 IBS=0.1096 t=0.041s
============================================================
Dataset: KidneyTransplant
============================================================
KidneyTransplant: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.4981 IBS=0.1435 t=0.049s
✓ AdaBoostRegressor C-index=0.6524 IBS=0.1451 t=0.132s
✓ BaggingRegressor C-index=0.6592 IBS=0.1769 t=0.156s
✓ BayesianRidge C-index=0.6947 IBS=0.1399 t=0.026s
✓ DecisionTreeRegressor C-index=0.6436 IBS=0.19 t=0.025s
✓ DummyRegressor C-index=0.5 IBS=0.1435 t=0.021s
✓ ElasticNet C-index=0.694 IBS=0.1411 t=0.024s
✓ ElasticNetCV C-index=0.694 IBS=0.1398 t=0.272s
✓ ExtraTreeRegressor C-index=0.6461 IBS=0.191 t=0.023s
✓ ExtraTreesRegressor C-index=0.6639 IBS=0.1898 t=1.051s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.651 IBS=0.1904 t=1.211s
✓ GradientBoostingRegressor C-index=0.68 IBS=0.1656 t=0.497s
✓ HistGradientBoostingRegressor C-index=0.6633 IBS=0.144 t=0.712s
✓ HuberRegressor C-index=0.6952 IBS=0.1421 t=0.113s
✓ KNeighborsRegressor C-index=0.6607 IBS=0.1486 t=0.035s
✓ KernelRidge C-index=0.6967 IBS=0.1398 t=0.164s
✓ Lars C-index=0.6991 IBS=0.1398 t=0.026s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.694 IBS=0.1427 t=0.02s
✓ LassoCV C-index=0.694 IBS=0.1398 t=0.279s
✓ LassoLars C-index=0.694 IBS=0.1427 t=0.024s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.694 IBS=0.1398 t=0.026s
✓ LinearRegression C-index=0.6991 IBS=0.1398 t=0.024s
✓ LinearSVR C-index=0.6744 IBS=0.1422 t=0.135s
✓ MLPRegressor C-index=0.6894 IBS=0.1407 t=0.734s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.6972 IBS=0.1422 t=0.183s
✓ OrthogonalMatchingPursuit C-index=0.694 IBS=0.1398 t=0.023s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.6952 IBS=0.1412 t=0.021s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.694 IBS=0.143 t=0.191s
✓ RANSACRegressor C-index=0.696 IBS=0.1414 t=0.544s
✓ RandomForestRegressor C-index=0.6556 IBS=0.1602 t=0.927s
✓ Ridge C-index=0.6971 IBS=0.1398 t=0.026s
✓ RidgeCV C-index=0.6982 IBS=0.1398 t=0.028s
✓ SGDRegressor C-index=0.5536 IBS=0.5984 t=0.041s
✓ SVR C-index=0.6965 IBS=0.1419 t=0.135s
✓ TheilSenRegressor C-index=0.6952 IBS=0.141 t=2.283s
✓ TweedieRegressor C-index=0.6947 IBS=0.1398 t=0.189s
============================================================
Dataset: NCTCGLung
============================================================
NCTCGLung: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.8904 IBS=nan t=0.051s
✓ AdaBoostRegressor C-index=0.3421 IBS=nan t=0.94s
✓ BaggingRegressor C-index=0.5088 IBS=nan t=0.359s
✓ BayesianRidge C-index=0.6667 IBS=nan t=0.027s
✓ DecisionTreeRegressor C-index=0.4868 IBS=nan t=0.015s
✓ DummyRegressor C-index=0.5 IBS=nan t=0.007s
✓ ElasticNet C-index=0.693 IBS=nan t=0.021s
✓ ElasticNetCV C-index=0.5 IBS=nan t=0.452s
✓ ExtraTreeRegressor C-index=0.4912 IBS=nan t=0.019s
✓ ExtraTreesRegressor C-index=0.6842 IBS=nan t=1.14s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.5 IBS=nan t=0.039s
✓ GradientBoostingRegressor C-index=0.5 IBS=nan t=0.595s
✓ HistGradientBoostingRegressor C-index=0.5 IBS=nan t=0.454s
✓ HuberRegressor C-index=0.5263 IBS=nan t=0.3s
✓ KNeighborsRegressor C-index=0.614 IBS=nan t=0.034s
✓ KernelRidge C-index=0.4825 IBS=nan t=0.017s
✓ Lars C-index=0.4561 IBS=nan t=0.042s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.5877 IBS=nan t=0.025s
✓ LassoCV C-index=0.5 IBS=nan t=0.479s
✓ LassoLars C-index=0.5877 IBS=nan t=0.026s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.5 IBS=nan t=0.028s
✓ LinearRegression C-index=0.4561 IBS=nan t=0.022s
✓ LinearSVR C-index=0.6053 IBS=nan t=0.066s
✓ MLPRegressor C-index=0.4386 IBS=nan t=0.605s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.4737 IBS=nan t=0.025s
✓ OrthogonalMatchingPursuit C-index=0.5877 IBS=nan t=0.01s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.3772 IBS=nan t=0.021s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.5921 IBS=nan t=0.09s
✓ RANSACRegressor C-index=0.2895 IBS=nan t=1.059s
✓ RandomForestRegressor C-index=0.5614 IBS=nan t=1.426s
✓ Ridge C-index=0.4649 IBS=nan t=0.013s
✓ RidgeCV C-index=0.5088 IBS=nan t=0.022s
✓ SGDRegressor C-index=0.4737 IBS=nan t=0.014s
✓ SVR C-index=0.6053 IBS=nan t=0.024s
✓ TheilSenRegressor C-index=0.4298 IBS=nan t=4.897s
✓ TweedieRegressor C-index=0.5877 IBS=nan t=0.147s
============================================================
Dataset: LymphNode
============================================================
LymphNode: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.6097 IBS=0.2348 t=0.055s
✓ AdaBoostRegressor C-index=0.688 IBS=0.1751 t=0.375s
✓ BaggingRegressor C-index=0.6695 IBS=0.2236 t=0.373s
✓ BayesianRidge C-index=0.6891 IBS=0.2376 t=0.039s
✓ DecisionTreeRegressor C-index=0.5334 IBS=0.2382 t=0.046s
✓ DummyRegressor C-index=0.5 IBS=0.1733 t=0.017s
✓ ElasticNet C-index=0.69 IBS=0.1731 t=0.023s
✓ ElasticNetCV C-index=0.6605 IBS=0.2364 t=0.484s
✓ ExtraTreeRegressor C-index=0.5017 IBS=0.2466 t=0.036s
✓ ExtraTreesRegressor C-index=0.6331 IBS=0.2321 t=2.121s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.5296 IBS=0.2495 t=0.474s
✓ GradientBoostingRegressor C-index=0.6539 IBS=0.2168 t=1.003s
✓ HistGradientBoostingRegressor C-index=0.6615 IBS=0.2137 t=1.162s
✓ HuberRegressor C-index=0.6789 IBS=0.2274 t=0.358s
✓ KNeighborsRegressor C-index=0.6666 IBS=0.1876 t=0.05s
✓ KernelRidge C-index=0.655 IBS=0.2337 t=0.422s
✓ Lars C-index=0.6472 IBS=0.2341 t=0.051s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.6965 IBS=0.1702 t=0.044s
✓ LassoCV C-index=0.6592 IBS=0.2357 t=0.665s
✓ LassoLars C-index=0.6965 IBS=0.1702 t=0.042s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.6495 IBS=0.2341 t=0.056s
✓ LinearRegression C-index=0.6472 IBS=0.2341 t=0.041s
✓ LinearSVR C-index=0.5184 IBS=0.8111 t=0.404s
✓ MLPRegressor C-index=0.5747 IBS=0.2844 t=3.176s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.6976 IBS=0.1808 t=0.238s
✓ OrthogonalMatchingPursuit C-index=0.7126 IBS=0.1702 t=0.027s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.6295 IBS=0.2823 t=0.025s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.6918 IBS=0.1753 t=0.306s
✓ RANSACRegressor C-index=0.5108 IBS=0.1752 t=0.976s
✓ RandomForestRegressor C-index=0.6725 IBS=0.2028 t=2.879s
✓ Ridge C-index=0.6474 IBS=0.2342 t=0.029s
✓ RidgeCV C-index=0.6488 IBS=0.2351 t=0.03s
✓ SGDRegressor C-index=0.3461 IBS=0.8417 t=0.037s
✓ SVR C-index=0.7052 IBS=0.1895 t=0.246s
✓ TheilSenRegressor C-index=0.6768 IBS=0.3194 t=5.491s
✓ TweedieRegressor C-index=0.6822 IBS=0.2421 t=0.176s
============================================================
Dataset: Leukemia
============================================================
Leukemia: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.8667 IBS=0.2412 t=0.024s
✓ AdaBoostRegressor C-index=0.8667 IBS=0.1425 t=0.328s
✓ BaggingRegressor C-index=0.8889 IBS=0.1551 t=0.096s
✓ BayesianRidge C-index=0.8667 IBS=0.2383 t=0.01s
✓ DecisionTreeRegressor C-index=0.8222 IBS=0.2156 t=0.01s
✓ DummyRegressor C-index=0.5 IBS=0.1957 t=0.004s
✓ ElasticNet C-index=0.5 IBS=0.1957 t=0.006s
✓ ElasticNetCV C-index=0.8667 IBS=0.2391 t=0.199s
✓ ExtraTreeRegressor C-index=0.8556 IBS=0.145 t=0.008s
✓ ExtraTreesRegressor C-index=0.8444 IBS=0.1446 t=0.418s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.5778 IBS=0.2871 t=0.014s
✓ GradientBoostingRegressor C-index=0.8444 IBS=0.194 t=0.225s
✓ HistGradientBoostingRegressor C-index=0.5 IBS=0.1957 t=0.113s
✓ HuberRegressor C-index=0.8667 IBS=0.2375 t=0.046s
✓ KNeighborsRegressor C-index=0.9556 IBS=0.1541 t=0.011s
✓ KernelRidge C-index=0.8 IBS=0.2091 t=0.021s
✓ Lars C-index=0.8667 IBS=0.2383 t=0.01s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.5 IBS=0.1957 t=0.009s
✓ LassoCV C-index=0.8667 IBS=0.2416 t=0.207s
✓ LassoLars C-index=0.5 IBS=0.1957 t=0.008s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.8667 IBS=0.2383 t=0.02s
✓ LinearRegression C-index=0.8667 IBS=0.2383 t=0.01s
✓ LinearSVR C-index=0.8667 IBS=0.2253 t=0.015s
✓ MLPRegressor C-index=0.8444 IBS=0.2026 t=0.318s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.8667 IBS=0.1493 t=0.008s
✓ OrthogonalMatchingPursuit C-index=0.8889 IBS=0.2616 t=0.007s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.8667 IBS=0.3682 t=0.006s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.5 IBS=0.1962 t=0.023s
✓ RANSACRegressor C-index=0.8667 IBS=0.2337 t=0.124s
✓ RandomForestRegressor C-index=0.9111 IBS=0.1719 t=0.532s
✓ Ridge C-index=0.8667 IBS=0.2381 t=0.008s
✓ RidgeCV C-index=0.8667 IBS=0.2381 t=0.009s
✓ SGDRegressor C-index=0.8 IBS=0.1878 t=0.01s
✓ SVR C-index=0.8667 IBS=0.1699 t=0.006s
✓ TheilSenRegressor C-index=0.8667 IBS=0.2423 t=1.129s
✓ TweedieRegressor C-index=0.9111 IBS=0.1817 t=0.035s
============================================================
Dataset: Larynx
============================================================
Larynx: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.5442 IBS=0.3101 t=0.037s
✓ AdaBoostRegressor C-index=0.6133 IBS=0.3205 t=0.112s
✓ BaggingRegressor C-index=0.6326 IBS=0.3216 t=0.129s
✓ BayesianRidge C-index=0.5608 IBS=0.3167 t=0.022s
✓ DecisionTreeRegressor C-index=0.6409 IBS=0.3313 t=0.007s
✓ DummyRegressor C-index=0.5 IBS=0.2469 t=0.004s
✓ ElasticNet C-index=0.558 IBS=0.2505 t=0.009s
✓ ElasticNetCV C-index=0.5608 IBS=0.329 t=0.284s
✓ ExtraTreeRegressor C-index=0.5912 IBS=0.3951 t=0.011s
✓ ExtraTreesRegressor C-index=0.6547 IBS=0.3533 t=0.509s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.5663 IBS=0.3326 t=0.02s
✓ GradientBoostingRegressor C-index=0.5221 IBS=0.3477 t=0.287s
✓ HistGradientBoostingRegressor C-index=0.6243 IBS=0.2521 t=0.181s
✓ HuberRegressor C-index=0.5663 IBS=0.3739 t=0.126s
✓ KNeighborsRegressor C-index=0.4392 IBS=0.3009 t=0.031s
✓ KernelRidge C-index=0.5663 IBS=0.3204 t=0.011s
✓ Lars C-index=0.5663 IBS=0.3464 t=0.014s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.558 IBS=0.2473 t=0.008s
✓ LassoCV C-index=0.5608 IBS=0.3305 t=0.349s
✓ LassoLars C-index=0.558 IBS=0.2473 t=0.016s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.5608 IBS=0.3359 t=0.02s
✓ LinearRegression C-index=0.5663 IBS=0.3464 t=0.016s
✓ LinearSVR C-index=0.511 IBS=0.2545 t=0.03s
✓ MLPRegressor C-index=0.5331 IBS=0.3079 t=0.327s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.5497 IBS=0.2589 t=0.014s
✓ OrthogonalMatchingPursuit C-index=0.558 IBS=0.2567 t=0.008s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.5497 IBS=0.3836 t=0.011s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.558 IBS=0.2767 t=0.051s
✓ RANSACRegressor C-index=0.5718 IBS=0.447 t=0.384s
✓ RandomForestRegressor C-index=0.6215 IBS=0.3291 t=1.13s
✓ Ridge C-index=0.5663 IBS=0.3335 t=0.014s
✓ RidgeCV C-index=0.5663 IBS=0.3335 t=0.017s
✓ SGDRegressor C-index=0.4392 IBS=0.4587 t=0.008s
✓ SVR C-index=0.5552 IBS=0.2978 t=0.009s
✓ TheilSenRegressor C-index=0.5718 IBS=0.2889 t=1.501s
✓ TweedieRegressor C-index=0.5718 IBS=0.2609 t=0.035s
============================================================
Dataset: Lymphoma
============================================================
Lymphoma: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.5549 IBS=0.1946 t=0.023s
✓ AdaBoostRegressor C-index=0.5549 IBS=0.1903 t=0.021s
✓ BaggingRegressor C-index=0.5549 IBS=0.1934 t=0.061s
✓ BayesianRidge C-index=0.5549 IBS=0.1946 t=0.006s
✓ DecisionTreeRegressor C-index=0.5549 IBS=0.1931 t=0.005s
✓ DummyRegressor C-index=0.5 IBS=0.2149 t=0.004s
✓ ElasticNet C-index=0.5 IBS=0.2149 t=0.005s
✓ ElasticNetCV C-index=0.5549 IBS=0.1936 t=0.122s
✓ ExtraTreeRegressor C-index=0.5549 IBS=0.1931 t=0.006s
✓ ExtraTreesRegressor C-index=0.5549 IBS=0.1931 t=0.208s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.5549 IBS=0.1931 t=0.01s
✓ GradientBoostingRegressor C-index=0.5549 IBS=0.1931 t=0.106s
✓ HistGradientBoostingRegressor C-index=0.5 IBS=0.2149 t=0.074s
✓ HuberRegressor C-index=0.5549 IBS=0.191 t=0.017s
✓ KNeighborsRegressor C-index=0.5549 IBS=0.1942 t=0.021s
✓ KernelRidge C-index=0.5549 IBS=0.1965 t=0.007s
✓ Lars C-index=0.5549 IBS=0.1931 t=0.005s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.5 IBS=0.2149 t=0.009s
✓ LassoCV C-index=0.5549 IBS=0.1933 t=0.111s
✓ LassoLars C-index=0.5 IBS=0.2149 t=0.005s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.5549 IBS=0.1931 t=0.008s
✓ LinearRegression C-index=0.5549 IBS=0.1931 t=0.007s
✓ LinearSVR C-index=0.5549 IBS=0.1945 t=0.006s
✓ MLPRegressor C-index=0.5549 IBS=0.1935 t=0.101s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.5549 IBS=0.191 t=0.007s
✓ OrthogonalMatchingPursuit C-index=0.5549 IBS=0.1931 t=0.004s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.5549 IBS=0.3721 t=0.004s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.5 IBS=0.2127 t=0.014s
✓ RANSACRegressor C-index=0.5549 IBS=0.2034 t=0.044s
✓ RandomForestRegressor C-index=0.5549 IBS=0.1929 t=0.319s
✓ Ridge C-index=0.5549 IBS=0.194 t=0.005s
✓ RidgeCV C-index=0.5549 IBS=0.194 t=0.007s
✓ SGDRegressor C-index=0.5549 IBS=0.2091 t=0.008s
✓ SVR C-index=0.5549 IBS=0.1924 t=0.008s
✓ TheilSenRegressor C-index=0.5549 IBS=0.2076 t=0.512s
✓ TweedieRegressor C-index=0.5549 IBS=0.2099 t=0.014s
============================================================
Dataset: StanfordHeart
============================================================
StanfordHeart: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.5865 IBS=0.2212 t=0.014s
✓ AdaBoostRegressor C-index=0.4549 IBS=0.2772 t=0.063s
✓ BaggingRegressor C-index=0.4211 IBS=0.3021 t=0.076s
✓ BayesianRidge C-index=0.5865 IBS=0.2212 t=0.005s
✓ DecisionTreeRegressor C-index=0.4361 IBS=0.3202 t=0.008s
✓ DummyRegressor C-index=0.5 IBS=0.2446 t=0.004s
✓ ElasticNet C-index=0.5865 IBS=0.2212 t=0.009s
✓ ElasticNetCV C-index=0.5865 IBS=0.2217 t=0.126s
✓ ExtraTreeRegressor C-index=0.5451 IBS=0.3094 t=0.005s
✓ ExtraTreesRegressor C-index=0.3985 IBS=0.3109 t=0.234s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.3083 IBS=0.5281 t=0.012s
✓ GradientBoostingRegressor C-index=0.4286 IBS=0.3073 t=0.113s
✓ HistGradientBoostingRegressor C-index=0.5301 IBS=0.2378 t=0.085s
✓ HuberRegressor C-index=0.5865 IBS=0.2253 t=0.015s
✓ KNeighborsRegressor C-index=0.5301 IBS=0.254 t=0.01s
✓ KernelRidge C-index=0.5865 IBS=0.2213 t=0.014s
✓ Lars C-index=0.5865 IBS=0.2228 t=0.006s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.5865 IBS=0.225 t=0.008s
✓ LassoCV C-index=0.5865 IBS=0.2217 t=0.114s
✓ LassoLars C-index=0.5865 IBS=0.225 t=0.007s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.5865 IBS=0.2228 t=0.01s
✓ LinearRegression C-index=0.5865 IBS=0.2228 t=0.004s
✓ LinearSVR C-index=0.4135 IBS=0.4936 t=0.012s
✓ MLPRegressor C-index=0.4135 IBS=0.3542 t=0.032s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.5865 IBS=0.2193 t=0.004s
✓ OrthogonalMatchingPursuit C-index=0.5865 IBS=0.2228 t=0.008s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.5865 IBS=0.2725 t=0.006s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.5865 IBS=0.2245 t=0.015s
✓ RANSACRegressor C-index=0.5865 IBS=0.2762 t=0.064s
✓ RandomForestRegressor C-index=0.4586 IBS=0.2942 t=0.28s
✓ Ridge C-index=0.5865 IBS=0.2228 t=0.006s
✓ RidgeCV C-index=0.5865 IBS=0.2228 t=0.005s
✓ SGDRegressor C-index=0.4135 IBS=0.8219 t=0.006s
✓ SVR C-index=0.5865 IBS=0.2442 t=0.005s
✓ TheilSenRegressor C-index=0.5865 IBS=0.23 t=0.564s
✓ TweedieRegressor C-index=0.5865 IBS=0.2226 t=0.013s
============================================================
Dataset: DemocracyDuration
============================================================
DemocracyDuration: 0%| | 0/45 [00:00<?, ?it/s]
✓ ARDRegression C-index=0.5863 IBS=0.0948 t=0.224s
✓ AdaBoostRegressor C-index=0.588 IBS=0.097 t=0.336s
✓ BaggingRegressor C-index=0.5968 IBS=0.1057 t=0.423s
✓ BayesianRidge C-index=0.5975 IBS=0.0981 t=0.074s
✓ DecisionTreeRegressor C-index=0.5954 IBS=0.1017 t=0.065s
✓ DummyRegressor C-index=0.5 IBS=0.1066 t=0.074s
✓ ElasticNet C-index=0.5113 IBS=0.1054 t=0.068s
✓ ElasticNetCV C-index=0.5982 IBS=0.098 t=0.953s
✓ ExtraTreeRegressor C-index=0.5997 IBS=0.101 t=0.06s
✓ ExtraTreesRegressor C-index=0.5987 IBS=0.1014 t=1.952s
✗ GammaRegressor C-index=nan IBS=nan t=nans
✓ GaussianProcessRegressor C-index=0.5901 IBS=0.102 t=3.655s
✓ GradientBoostingRegressor C-index=0.6005 IBS=0.0995 t=1.123s
✓ HistGradientBoostingRegressor C-index=0.614 IBS=0.1036 t=2.613s
✓ HuberRegressor C-index=0.6046 IBS=0.0982 t=1.644s
✓ KNeighborsRegressor C-index=0.611 IBS=0.1071 t=0.121s
✓ KernelRidge C-index=0.5876 IBS=0.0956 t=1.681s
✓ Lars C-index=0.6018 IBS=0.0981 t=0.061s
✗ LarsCV C-index=nan IBS=nan t=nans
✓ Lasso C-index=0.5113 IBS=0.1065 t=0.044s
✓ LassoCV C-index=0.5982 IBS=0.0978 t=0.578s
✓ LassoLars C-index=0.5113 IBS=0.1065 t=0.051s
✗ LassoLarsCV C-index=nan IBS=nan t=nans
✓ LassoLarsIC C-index=0.5977 IBS=0.0978 t=0.092s
✓ LinearRegression C-index=0.5977 IBS=0.0978 t=0.068s
✓ LinearSVR C-index=0.5817 IBS=0.1143 t=0.562s
✓ MLPRegressor C-index=0.5893 IBS=0.1026 t=8.281s
✗ MultiTaskElasticNet C-index=nan IBS=nan t=nans
✗ MultiTaskElasticNetCV C-index=nan IBS=nan t=nans
✗ MultiTaskLasso C-index=nan IBS=nan t=nans
✗ MultiTaskLassoCV C-index=nan IBS=nan t=nans
✓ NuSVR C-index=0.5199 IBS=0.1054 t=1.336s
✓ OrthogonalMatchingPursuit C-index=0.5113 IBS=0.1046 t=0.048s
✗ OrthogonalMatchingPursuitCV C-index=nan IBS=nan t=nans
✓ PassiveAggressiveRegressor C-index=0.5182 IBS=0.1128 t=0.054s
✗ PoissonRegressor C-index=nan IBS=nan t=nans
✓ QuantileRegressor C-index=0.5 IBS=0.1057 t=1.006s
✓ RANSACRegressor C-index=0.5566 IBS=0.1073 t=1.149s
✓ RandomForestRegressor C-index=0.6032 IBS=0.1041 t=2.841s
✓ Ridge C-index=0.5975 IBS=0.0979 t=0.071s
✓ RidgeCV C-index=0.5975 IBS=0.0979 t=0.211s
✓ SGDRegressor C-index=0.4871 IBS=0.924 t=0.145s
✓ SVR C-index=0.5536 IBS=0.1064 t=1.983s
✓ TheilSenRegressor C-index=0.5979 IBS=0.0984 t=6.926s
✓ TweedieRegressor C-index=0.5803 IBS=0.1007 t=0.214s
Total experiments: 675
Successful: 540
Failed: 135
── Baselines: WHAS500 ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.7191 IBS=0.1958 t=0.075s
✓ sksurv/RandomSurvivalForest C-index=0.7525 IBS=0.1788 t=0.704s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.7392 IBS=0.205 t=0.483s
✓ lifelines/CoxPHFitter C-index=0.7273 IBS=nan t=0.222s
── Baselines: GBSG2 ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.6441 IBS=0.1731 t=0.07s
✓ sksurv/RandomSurvivalForest C-index=0.6713 IBS=0.1671 t=0.716s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.6414 IBS=0.1753 t=0.53s
✓ lifelines/CoxPHFitter C-index=0.6641 IBS=nan t=0.107s
── Baselines: VeteransLungCancer ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.6696 IBS=nan t=0.015s
✓ sksurv/RandomSurvivalForest C-index=0.633 IBS=nan t=0.133s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.687 IBS=nan t=0.073s
✓ lifelines/CoxPHFitter C-index=0.6661 IBS=nan t=0.053s
── Baselines: FLChain ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.9322 IBS=0.0436 t=1.214s
✓ sksurv/RandomSurvivalForest C-index=0.9316 IBS=0.048 t=33.408s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.9242 IBS=0.0536 t=61.798s
✓ lifelines/CoxPHFitter C-index=0.9321 IBS=nan t=1.963s
── Baselines: AIDS ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.7718 IBS=0.0654 t=0.069s
✓ sksurv/RandomSurvivalForest C-index=0.7241 IBS=0.0685 t=0.475s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.7244 IBS=0.0718 t=1.141s
✓ lifelines/CoxPHFitter C-index=0.7554 IBS=nan t=0.11s
── Baselines: BreastCancerGenomic ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.6648 IBS=0.9192 t=0.079s
✓ sksurv/RandomSurvivalForest C-index=0.651 IBS=0.2311 t=0.5s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.6786 IBS=0.2285 t=0.373s
✗ lifelines/CoxPHFitter C-index=nan IBS=nan t=nans
── Baselines: Rossi ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.5646 IBS=0.1064 t=0.023s
✓ sksurv/RandomSurvivalForest C-index=0.6517 IBS=0.1035 t=0.134s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.6213 IBS=0.1064 t=0.197s
✓ lifelines/CoxPHFitter C-index=0.5856 IBS=nan t=0.048s
── Baselines: KidneyTransplant ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.6987 IBS=0.1261 t=0.041s
✓ sksurv/RandomSurvivalForest C-index=0.6426 IBS=0.1437 t=0.499s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.6648 IBS=0.1301 t=0.644s
✓ lifelines/CoxPHFitter C-index=0.6967 IBS=nan t=0.053s
── Baselines: NCTCGLung ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.4474 IBS=nan t=0.012s
✓ sksurv/RandomSurvivalForest C-index=0.4035 IBS=nan t=0.131s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.4825 IBS=nan t=0.082s
✓ lifelines/CoxPHFitter C-index=0.6228 IBS=nan t=0.042s
── Baselines: LymphNode ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.6817 IBS=0.1458 t=0.071s
✓ sksurv/RandomSurvivalForest C-index=0.6568 IBS=0.1499 t=0.569s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.6964 IBS=0.1486 t=0.493s
✓ lifelines/CoxPHFitter C-index=0.6564 IBS=nan t=0.072s
── Baselines: Leukemia ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.9111 IBS=0.0904 t=0.006s
✓ sksurv/RandomSurvivalForest C-index=0.9556 IBS=0.0784 t=0.105s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.8889 IBS=0.094 t=0.052s
✓ lifelines/CoxPHFitter C-index=0.8667 IBS=nan t=0.036s
── Baselines: Larynx ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.5663 IBS=0.246 t=0.008s
✓ sksurv/RandomSurvivalForest C-index=0.5718 IBS=0.2278 t=0.11s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.5193 IBS=0.2897 t=0.057s
✓ lifelines/CoxPHFitter C-index=0.5718 IBS=nan t=0.045s
── Baselines: Lymphoma ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.5549 IBS=0.1956 t=0.009s
✓ sksurv/RandomSurvivalForest C-index=0.5549 IBS=0.1955 t=0.12s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.5549 IBS=0.1956 t=0.051s
✓ lifelines/CoxPHFitter C-index=0.5549 IBS=nan t=0.035s
── Baselines: StanfordHeart ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.5865 IBS=0.2196 t=0.008s
✓ sksurv/RandomSurvivalForest C-index=0.5639 IBS=0.302 t=0.11s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.4098 IBS=0.3776 t=0.059s
✓ lifelines/CoxPHFitter C-index=0.5865 IBS=nan t=0.031s
── Baselines: DemocracyDuration ──
✓ sksurv/CoxPHSurvivalAnalysis C-index=0.5946 IBS=0.0998 t=0.066s
✓ sksurv/RandomSurvivalForest C-index=0.5891 IBS=0.121 t=0.508s
✓ sksurv/GradientBoostingSurvivalAnalysis C-index=0.6113 IBS=0.0915 t=3.601s
✓ lifelines/CoxPHFitter C-index=0.5916 IBS=nan t=0.067s
✓ baselines done
Saved 735 rows to survival_benchmark_results.csv
════════════════════════════════════════════════════════════════════════════════════════════════════
WHAS500 — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
sksurv RandomSurvivalForest 0.7525 0.1578 0.1788 0.704
survivalist SurvivalCustom(ExtraTreesRegressor) 0.7462 0.3436 0.3434 6.299
sksurv GradientBoostingSurvivalAnalysis 0.7392 0.1820 0.2050 0.483
lifelines CoxPHFitter 0.7273 NaN NaN 0.222
survivalist SurvivalCustom(LinearSVR) 0.7246 0.3838 0.3753 0.587
survivalist SurvivalCustom(KernelRidge) 0.7208 0.3159 0.3152 0.433
survivalist SurvivalCustom(Lars) 0.7199 0.3149 0.3143 0.114
survivalist SurvivalCustom(HuberRegressor) 0.7193 0.3420 0.3390 0.952
survivalist SurvivalCustom(Ridge) 0.7191 0.3151 0.3145 0.043
sksurv CoxPHSurvivalAnalysis 0.7191 0.1696 0.1958 0.075
survivalist SurvivalCustom(LinearRegression) 0.7189 0.3168 0.3160 0.044
survivalist SurvivalCustom(RidgeCV) 0.7187 0.3120 0.3117 0.095
survivalist SurvivalCustom(LassoLarsIC) 0.7174 0.3047 0.3048 0.114
survivalist SurvivalCustom(TheilSenRegressor) 0.7174 0.3147 0.3144 26.558
survivalist SurvivalCustom(ARDRegression) 0.7170 0.3026 0.3011 0.148
survivalist SurvivalCustom(LassoCV) 0.7168 0.3072 0.3072 1.225
survivalist SurvivalCustom(ElasticNetCV) 0.7163 0.3073 0.3072 1.359
survivalist SurvivalCustom(RandomForestRegressor) 0.7140 0.3269 0.3285 8.702
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.7072 0.4576 0.4363 0.046
survivalist SurvivalCustom(GradientBoostingRegressor) 0.6986 0.3369 0.3380 4.199
survivalist SurvivalCustom(TweedieRegressor) 0.6962 0.3151 0.3129 0.477
survivalist SurvivalCustom(RANSACRegressor) 0.6876 0.3960 0.3900 2.657
survivalist SurvivalCustom(BayesianRidge) 0.6873 0.3258 0.3218 0.088
survivalist SurvivalCustom(NuSVR) 0.6846 0.2811 0.2810 0.287
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.6826 0.2757 0.2770 0.029
survivalist SurvivalCustom(AdaBoostRegressor) 0.6819 0.3073 0.3025 1.809
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.6818 0.3916 0.3788 2.730
survivalist SurvivalCustom(QuantileRegressor) 0.6810 0.3258 0.3223 0.650
survivalist SurvivalCustom(SVR) 0.6806 0.3622 0.3555 0.406
survivalist SurvivalCustom(LassoLars) 0.6795 0.2825 0.2815 0.050
survivalist SurvivalCustom(Lasso) 0.6795 0.2825 0.2815 0.036
survivalist SurvivalCustom(ElasticNet) 0.6787 0.3055 0.3030 0.036
survivalist SurvivalCustom(KNeighborsRegressor) 0.6774 0.3117 0.3095 0.059
survivalist SurvivalCustom(BaggingRegressor) 0.6725 0.3517 0.3496 1.583
survivalist SurvivalCustom(MLPRegressor) 0.6431 0.4274 0.4129 7.804
survivalist SurvivalCustom(DecisionTreeRegressor) 0.6251 0.3453 0.3495 0.128
survivalist SurvivalCustom(ExtraTreeRegressor) 0.6242 0.3664 0.3630 0.069
survivalist SurvivalCustom(GaussianProcessRegressor) 0.5005 0.4029 0.3858 1.071
survivalist SurvivalCustom(DummyRegressor) 0.5000 0.2404 0.2338 0.018
survivalist SurvivalCustom(SGDRegressor) 0.4395 0.6485 0.6711 0.131
════════════════════════════════════════════════════════════════════════════════════════════════════
GBSG2 — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
survivalist SurvivalCustom(ARDRegression) 0.7098 0.4496 0.3462 0.076
survivalist SurvivalCustom(HuberRegressor) 0.7020 0.5019 0.3873 0.496
survivalist SurvivalCustom(KernelRidge) 0.6840 0.4392 0.3387 0.460
survivalist SurvivalCustom(LassoLarsIC) 0.6833 0.4388 0.3384 0.071
survivalist SurvivalCustom(Lars) 0.6833 0.4388 0.3384 0.046
survivalist SurvivalCustom(LinearRegression) 0.6833 0.4388 0.3384 0.029
survivalist SurvivalCustom(Ridge) 0.6833 0.4391 0.3386 0.050
survivalist SurvivalCustom(RidgeCV) 0.6832 0.4410 0.3399 0.052
survivalist SurvivalCustom(TheilSenRegressor) 0.6742 0.5459 0.4324 7.961
sksurv RandomSurvivalForest 0.6713 0.2213 0.1671 0.716
survivalist SurvivalCustom(TweedieRegressor) 0.6658 0.4415 0.3398 0.235
lifelines CoxPHFitter 0.6641 NaN NaN 0.107
survivalist SurvivalCustom(ElasticNet) 0.6565 0.2907 0.2330 0.028
survivalist SurvivalCustom(RANSACRegressor) 0.6523 0.5838 0.5078 1.412
survivalist SurvivalCustom(BayesianRidge) 0.6502 0.4089 0.3168 0.038
survivalist SurvivalCustom(ElasticNetCV) 0.6486 0.3965 0.3082 0.712
survivalist SurvivalCustom(LassoCV) 0.6475 0.3939 0.3064 0.680
survivalist SurvivalCustom(AdaBoostRegressor) 0.6447 0.2692 0.2121 0.750
sksurv CoxPHSurvivalAnalysis 0.6441 0.2230 0.1731 0.070
sksurv GradientBoostingSurvivalAnalysis 0.6414 0.2240 0.1753 0.530
survivalist SurvivalCustom(GradientBoostingRegressor) 0.6228 0.3271 0.2570 1.528
survivalist SurvivalCustom(ExtraTreesRegressor) 0.6221 0.3595 0.2790 4.611
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.6191 0.4216 0.3391 0.027
survivalist SurvivalCustom(Lasso) 0.6150 0.2430 0.1912 0.027
survivalist SurvivalCustom(LassoLars) 0.6150 0.2430 0.1912 0.031
survivalist SurvivalCustom(LinearSVR) 0.6137 0.5832 0.4824 0.456
survivalist SurvivalCustom(RandomForestRegressor) 0.6129 0.3353 0.2625 4.755
survivalist SurvivalCustom(BaggingRegressor) 0.6114 0.3506 0.2722 0.602
survivalist SurvivalCustom(QuantileRegressor) 0.6071 0.2447 0.1899 0.437
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.6071 0.2429 0.1899 0.025
survivalist SurvivalCustom(NuSVR) 0.6062 0.2598 0.2051 0.350
survivalist SurvivalCustom(SVR) 0.6044 0.2995 0.2354 0.761
survivalist SurvivalCustom(ExtraTreeRegressor) 0.5961 0.3730 0.2942 0.045
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.5952 0.3603 0.2776 1.806
survivalist SurvivalCustom(DecisionTreeRegressor) 0.5840 0.3561 0.2770 0.067
survivalist SurvivalCustom(MLPRegressor) 0.5739 0.5647 0.4639 5.809
survivalist SurvivalCustom(GaussianProcessRegressor) 0.5565 0.3859 0.3011 1.237
survivalist SurvivalCustom(KNeighborsRegressor) 0.5529 0.3507 0.2697 0.078
survivalist SurvivalCustom(SGDRegressor) 0.5508 0.4092 0.3983 0.063
survivalist SurvivalCustom(DummyRegressor) 0.5000 0.2460 0.1916 0.019
════════════════════════════════════════════════════════════════════════════════════════════════════
VeteransLungCancer — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
survivalist SurvivalCustom(ElasticNet) 0.7078 NaN NaN 0.036
survivalist SurvivalCustom(QuantileRegressor) 0.7078 NaN NaN 0.100
survivalist SurvivalCustom(Lasso) 0.7078 NaN NaN 0.014
survivalist SurvivalCustom(SVR) 0.7078 NaN NaN 0.025
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.7078 NaN NaN 0.011
survivalist SurvivalCustom(LassoLars) 0.7078 NaN NaN 0.017
survivalist SurvivalCustom(NuSVR) 0.7061 NaN NaN 0.038
survivalist SurvivalCustom(BayesianRidge) 0.7026 NaN NaN 0.084
survivalist SurvivalCustom(TweedieRegressor) 0.7009 NaN NaN 0.198
sksurv GradientBoostingSurvivalAnalysis 0.6870 NaN NaN 0.073
survivalist SurvivalCustom(RidgeCV) 0.6713 NaN NaN 0.027
sksurv CoxPHSurvivalAnalysis 0.6696 NaN NaN 0.015
lifelines CoxPHFitter 0.6661 NaN NaN 0.053
survivalist SurvivalCustom(HuberRegressor) 0.6574 NaN NaN 0.383
survivalist SurvivalCustom(ElasticNetCV) 0.6557 NaN NaN 0.896
survivalist SurvivalCustom(MLPRegressor) 0.6539 NaN NaN 0.619
survivalist SurvivalCustom(RandomForestRegressor) 0.6504 NaN NaN 1.800
survivalist SurvivalCustom(LassoCV) 0.6504 NaN NaN 0.646
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.6487 NaN NaN 0.531
survivalist SurvivalCustom(TheilSenRegressor) 0.6470 NaN NaN 7.139
survivalist SurvivalCustom(LassoLarsIC) 0.6452 NaN NaN 0.045
survivalist SurvivalCustom(KNeighborsRegressor) 0.6443 NaN NaN 0.037
survivalist SurvivalCustom(DecisionTreeRegressor) 0.6426 NaN NaN 0.029
survivalist SurvivalCustom(LinearRegression) 0.6383 NaN NaN 0.015
survivalist SurvivalCustom(Lars) 0.6383 NaN NaN 0.035
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.6365 NaN NaN 0.020
survivalist SurvivalCustom(Ridge) 0.6365 NaN NaN 0.016
survivalist SurvivalCustom(KernelRidge) 0.6330 NaN NaN 0.018
sksurv RandomSurvivalForest 0.6330 NaN NaN 0.133
survivalist SurvivalCustom(LinearSVR) 0.6313 NaN NaN 0.069
survivalist SurvivalCustom(ARDRegression) 0.6270 NaN NaN 0.125
survivalist SurvivalCustom(AdaBoostRegressor) 0.6243 NaN NaN 1.045
survivalist SurvivalCustom(ExtraTreesRegressor) 0.6209 NaN NaN 1.917
survivalist SurvivalCustom(ExtraTreeRegressor) 0.6070 NaN NaN 0.034
survivalist SurvivalCustom(BaggingRegressor) 0.6052 NaN NaN 0.461
survivalist SurvivalCustom(GradientBoostingRegressor) 0.6035 NaN NaN 0.761
survivalist SurvivalCustom(RANSACRegressor) 0.6000 NaN NaN 1.426
survivalist SurvivalCustom(SGDRegressor) 0.5617 NaN NaN 0.019
survivalist SurvivalCustom(GaussianProcessRegressor) 0.5087 NaN NaN 0.046
survivalist SurvivalCustom(DummyRegressor) 0.5000 NaN NaN 0.013
════════════════════════════════════════════════════════════════════════════════════════════════════
FLChain — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
sksurv CoxPHSurvivalAnalysis 0.9322 0.0650 0.0436 1.214
survivalist SurvivalCustom(ExtraTreesRegressor) 0.9322 0.1260 0.1185 184.103
lifelines CoxPHFitter 0.9321 NaN NaN 1.963
sksurv RandomSurvivalForest 0.9316 0.0679 0.0480 33.408
survivalist SurvivalCustom(KernelRidge) 0.9312 0.1354 0.1276 176.373
survivalist SurvivalCustom(Ridge) 0.9312 0.1358 0.1280 1.107
survivalist SurvivalCustom(RandomForestRegressor) 0.9309 0.1239 0.1158 225.531
survivalist SurvivalCustom(BaggingRegressor) 0.9304 0.1255 0.1176 24.055
survivalist SurvivalCustom(MLPRegressor) 0.9304 0.1447 0.1367 62.198
survivalist SurvivalCustom(RidgeCV) 0.9301 0.1287 0.1206 2.316
survivalist SurvivalCustom(LinearRegression) 0.9299 0.1286 0.1205 3.134
survivalist SurvivalCustom(LassoLarsIC) 0.9299 0.1286 0.1205 4.031
survivalist SurvivalCustom(BayesianRidge) 0.9299 0.1286 0.1205 1.498
survivalist SurvivalCustom(LinearSVR) 0.9289 0.1328 0.1247 19.291
survivalist SurvivalCustom(Lars) 0.9288 0.1505 0.1424 1.288
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.9283 0.1495 0.1424 5.929
survivalist SurvivalCustom(GradientBoostingRegressor) 0.9279 0.1218 0.1138 42.219
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.9274 0.1491 0.1414 26.807
survivalist SurvivalCustom(LassoCV) 0.9274 0.1741 0.1663 8.204
survivalist SurvivalCustom(ElasticNetCV) 0.9267 0.1852 0.1766 5.788
sksurv GradientBoostingSurvivalAnalysis 0.9242 0.0712 0.0536 61.798
survivalist SurvivalCustom(DecisionTreeRegressor) 0.9229 0.1266 0.1190 3.875
survivalist SurvivalCustom(ExtraTreeRegressor) 0.9229 0.1263 0.1190 2.492
survivalist SurvivalCustom(TheilSenRegressor) 0.9225 0.1429 0.1350 214.080
survivalist SurvivalCustom(ARDRegression) 0.9223 0.1283 0.1201 1.259
survivalist SurvivalCustom(HuberRegressor) 0.9193 0.2742 0.2541 17.314
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.8876 0.2383 0.2241 0.898
survivalist SurvivalCustom(AdaBoostRegressor) 0.8854 0.1362 0.1283 5.704
survivalist SurvivalCustom(GaussianProcessRegressor) 0.8761 0.1448 0.1352 556.617
survivalist SurvivalCustom(NuSVR) 0.8610 0.2826 0.2591 160.384
survivalist SurvivalCustom(SVR) 0.8599 0.2875 0.2630 149.160
survivalist SurvivalCustom(RANSACRegressor) 0.8328 0.1665 0.1574 12.349
survivalist SurvivalCustom(TweedieRegressor) 0.8094 0.1856 0.1760 1.138
survivalist SurvivalCustom(KNeighborsRegressor) 0.8074 0.1548 0.1462 13.201
survivalist SurvivalCustom(Lasso) 0.7722 0.1437 0.1329 0.851
survivalist SurvivalCustom(ElasticNet) 0.7722 0.1508 0.1413 1.199
survivalist SurvivalCustom(QuantileRegressor) 0.7722 0.1450 0.1300 75.045
survivalist SurvivalCustom(LassoLars) 0.7722 0.1437 0.1329 1.001
survivalist SurvivalCustom(SGDRegressor) 0.5146 0.8170 0.8268 8.477
survivalist SurvivalCustom(DummyRegressor) 0.5000 0.1452 0.1305 0.662
════════════════════════════════════════════════════════════════════════════════════════════════════
AIDS — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
sksurv CoxPHSurvivalAnalysis 0.7718 0.0744 0.0654 0.069
survivalist SurvivalCustom(ElasticNet) 0.7630 0.0842 0.0733 0.060
survivalist SurvivalCustom(LassoLars) 0.7630 0.0843 0.0734 0.080
survivalist SurvivalCustom(Lasso) 0.7630 0.0843 0.0734 0.053
survivalist SurvivalCustom(LassoCV) 0.7610 0.0834 0.0727 2.251
survivalist SurvivalCustom(ElasticNetCV) 0.7610 0.0835 0.0727 2.761
survivalist SurvivalCustom(BayesianRidge) 0.7600 0.0836 0.0728 0.158
lifelines CoxPHFitter 0.7554 NaN NaN 0.110
survivalist SurvivalCustom(LassoLarsIC) 0.7536 0.0831 0.0725 1.074
survivalist SurvivalCustom(TweedieRegressor) 0.7518 0.0838 0.0730 0.478
survivalist SurvivalCustom(AdaBoostRegressor) 0.7500 0.0882 0.0784 0.783
survivalist SurvivalCustom(RidgeCV) 0.7461 0.0831 0.0724 0.128
survivalist SurvivalCustom(KernelRidge) 0.7424 0.0830 0.0723 2.222
survivalist SurvivalCustom(Ridge) 0.7422 0.0830 0.0723 0.080
survivalist SurvivalCustom(LinearRegression) 0.7415 0.0830 0.0723 0.103
survivalist SurvivalCustom(TheilSenRegressor) 0.7399 0.0840 0.0732 42.094
survivalist SurvivalCustom(SGDRegressor) 0.7398 0.0971 0.0860 0.280
survivalist SurvivalCustom(ARDRegression) 0.7366 0.0832 0.0725 0.247
survivalist SurvivalCustom(Lars) 0.7360 0.0830 0.0724 0.228
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.7306 0.0839 0.0731 0.064
sksurv GradientBoostingSurvivalAnalysis 0.7244 0.0830 0.0718 1.141
sksurv RandomSurvivalForest 0.7241 0.0774 0.0685 0.475
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.7147 0.0830 0.0724 0.081
survivalist SurvivalCustom(MLPRegressor) 0.7139 0.0838 0.0736 27.167
survivalist SurvivalCustom(GradientBoostingRegressor) 0.6962 0.0865 0.0758 4.841
survivalist SurvivalCustom(RandomForestRegressor) 0.6726 0.0858 0.0752 17.432
survivalist SurvivalCustom(SVR) 0.6709 0.0847 0.0738 0.978
survivalist SurvivalCustom(LinearSVR) 0.6687 0.0831 0.0726 2.106
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.6587 0.0861 0.0756 8.208
survivalist SurvivalCustom(KNeighborsRegressor) 0.6527 0.0832 0.0726 0.212
survivalist SurvivalCustom(ExtraTreesRegressor) 0.6429 0.0903 0.0795 14.052
survivalist SurvivalCustom(BaggingRegressor) 0.6309 0.0904 0.0800 1.823
survivalist SurvivalCustom(HuberRegressor) 0.5720 0.0850 0.0740 1.691
survivalist SurvivalCustom(NuSVR) 0.5596 0.0850 0.0740 2.886
survivalist SurvivalCustom(RANSACRegressor) 0.5260 0.0849 0.0740 3.449
survivalist SurvivalCustom(DecisionTreeRegressor) 0.5148 0.0906 0.0798 0.242
survivalist SurvivalCustom(GaussianProcessRegressor) 0.5034 0.0901 0.0797 12.940
survivalist SurvivalCustom(DummyRegressor) 0.5000 0.0848 0.0739 0.040
survivalist SurvivalCustom(QuantileRegressor) 0.5000 0.0851 0.0741 1.840
survivalist SurvivalCustom(ExtraTreeRegressor) 0.4762 0.0987 0.0890 0.213
════════════════════════════════════════════════════════════════════════════════════════════════════
BreastCancerGenomic — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
sksurv GradientBoostingSurvivalAnalysis 0.6786 0.2464 0.2285 0.373
sksurv CoxPHSurvivalAnalysis 0.6648 0.7508 0.9192 0.079
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.6538 0.2256 0.2583 12.298
sksurv RandomSurvivalForest 0.6510 0.2472 0.2311 0.500
survivalist SurvivalCustom(ExtraTreesRegressor) 0.6497 0.2311 0.2630 31.125
survivalist SurvivalCustom(AdaBoostRegressor) 0.6476 0.2144 0.2305 18.053
survivalist SurvivalCustom(ARDRegression) 0.6455 0.2238 0.2353 7.201
survivalist SurvivalCustom(TweedieRegressor) 0.6441 0.2298 0.2274 1.736
survivalist SurvivalCustom(TheilSenRegressor) 0.6428 0.2621 0.3167 1618.689
survivalist SurvivalCustom(BayesianRidge) 0.6359 0.2379 0.2271 1.312
survivalist SurvivalCustom(LinearSVR) 0.6331 0.2353 0.2670 3.115
survivalist SurvivalCustom(RidgeCV) 0.6317 0.2309 0.2596 0.443
survivalist SurvivalCustom(SVR) 0.6317 0.2604 0.2351 0.281
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.6303 0.2507 0.2296 0.212
survivalist SurvivalCustom(HuberRegressor) 0.6221 0.2395 0.2731 3.563
survivalist SurvivalCustom(NuSVR) 0.6193 0.2629 0.2366 0.337
survivalist SurvivalCustom(KernelRidge) 0.6166 0.2542 0.3000 0.748
survivalist SurvivalCustom(Ridge) 0.6152 0.2562 0.3020 0.211
survivalist SurvivalCustom(LinearRegression) 0.6014 0.2666 0.3157 0.364
survivalist SurvivalCustom(Lars) 0.5959 0.2747 0.3238 1.879
survivalist SurvivalCustom(ElasticNetCV) 0.5903 0.2429 0.2308 26.208
survivalist SurvivalCustom(LassoCV) 0.5890 0.2425 0.2308 27.687
survivalist SurvivalCustom(MLPRegressor) 0.5862 0.2477 0.2658 16.889
survivalist SurvivalCustom(LassoLarsIC) 0.5834 0.2411 0.2309 4.653
survivalist SurvivalCustom(DecisionTreeRegressor) 0.5579 0.2593 0.3154 0.730
survivalist SurvivalCustom(GradientBoostingRegressor) 0.5393 0.2412 0.2844 33.544
survivalist SurvivalCustom(KNeighborsRegressor) 0.5317 0.2383 0.2407 0.131
survivalist SurvivalCustom(RandomForestRegressor) 0.5241 0.2380 0.2503 67.673
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.5214 0.2351 0.2509 0.234
survivalist SurvivalCustom(DummyRegressor) 0.5000 0.2645 0.2379 0.042
survivalist SurvivalCustom(ElasticNet) 0.5000 0.2645 0.2379 0.092
survivalist SurvivalCustom(QuantileRegressor) 0.5000 0.2764 0.2449 2.956
survivalist SurvivalCustom(LassoLars) 0.5000 0.2645 0.2379 0.445
survivalist SurvivalCustom(GaussianProcessRegressor) 0.5000 0.2455 0.2742 0.876
survivalist SurvivalCustom(Lasso) 0.5000 0.2645 0.2379 0.118
survivalist SurvivalCustom(ExtraTreeRegressor) 0.4703 0.2889 0.2868 0.338
survivalist SurvivalCustom(BaggingRegressor) 0.4579 0.2547 0.2860 7.901
survivalist SurvivalCustom(SGDRegressor) 0.4000 0.7508 0.9192 0.286
survivalist SurvivalCustom(RANSACRegressor) 0.3821 0.6669 0.7496 22.216
════════════════════════════════════════════════════════════════════════════════════════════════════
Rossi — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
survivalist SurvivalCustom(BayesianRidge) 0.6750 0.1141 0.1094 0.035
survivalist SurvivalCustom(TweedieRegressor) 0.6610 0.1146 0.1096 0.041
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.6603 0.1129 0.1105 0.022
sksurv RandomSurvivalForest 0.6517 0.1039 0.1035 0.134
survivalist SurvivalCustom(ElasticNetCV) 0.6240 0.1152 0.1100 0.438
sksurv GradientBoostingSurvivalAnalysis 0.6213 0.1079 0.1064 0.197
survivalist SurvivalCustom(GradientBoostingRegressor) 0.6186 0.1518 0.1346 0.587
survivalist SurvivalCustom(LassoCV) 0.6173 0.1152 0.1100 0.615
survivalist SurvivalCustom(SVR) 0.6143 0.1128 0.1124 0.114
survivalist SurvivalCustom(KNeighborsRegressor) 0.6121 0.1268 0.1149 0.042
survivalist SurvivalCustom(NuSVR) 0.6106 0.1129 0.1114 0.115
survivalist SurvivalCustom(RandomForestRegressor) 0.6054 0.1343 0.1237 1.335
survivalist SurvivalCustom(LassoLarsIC) 0.6036 0.1155 0.1102 0.046
survivalist SurvivalCustom(TheilSenRegressor) 0.6006 0.1131 0.1095 3.562
survivalist SurvivalCustom(BaggingRegressor) 0.5996 0.1392 0.1278 0.233
survivalist SurvivalCustom(AdaBoostRegressor) 0.5979 0.1265 0.1162 0.142
survivalist SurvivalCustom(GaussianProcessRegressor) 0.5978 0.1440 0.1317 0.128
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.5966 0.1254 0.1154 0.726
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.5873 0.1123 0.1101 0.023
lifelines CoxPHFitter 0.5856 NaN NaN 0.048
survivalist SurvivalCustom(HuberRegressor) 0.5836 0.1134 0.1137 0.403
survivalist SurvivalCustom(MLPRegressor) 0.5799 0.1528 0.1365 1.641
survivalist SurvivalCustom(RidgeCV) 0.5792 0.1170 0.1111 0.022
survivalist SurvivalCustom(Ridge) 0.5706 0.1175 0.1115 0.014
survivalist SurvivalCustom(DecisionTreeRegressor) 0.5699 0.1608 0.1441 0.020
survivalist SurvivalCustom(KernelRidge) 0.5689 0.1175 0.1115 0.066
survivalist SurvivalCustom(ExtraTreesRegressor) 0.5689 0.1500 0.1368 1.135
survivalist SurvivalCustom(Lars) 0.5686 0.1175 0.1115 0.031
survivalist SurvivalCustom(LinearRegression) 0.5686 0.1175 0.1115 0.021
sksurv CoxPHSurvivalAnalysis 0.5646 0.1073 0.1064 0.023
survivalist SurvivalCustom(LinearSVR) 0.5619 0.1121 0.1088 0.186
survivalist SurvivalCustom(ARDRegression) 0.5559 0.1151 0.1108 0.050
survivalist SurvivalCustom(ExtraTreeRegressor) 0.5516 0.1656 0.1506 0.019
survivalist SurvivalCustom(SGDRegressor) 0.5379 0.2407 0.2751 0.085
survivalist SurvivalCustom(ElasticNet) 0.5000 0.1129 0.1117 0.018
survivalist SurvivalCustom(DummyRegressor) 0.5000 0.1129 0.1117 0.007
survivalist SurvivalCustom(Lasso) 0.5000 0.1129 0.1117 0.021
survivalist SurvivalCustom(LassoLars) 0.5000 0.1129 0.1117 0.027
survivalist SurvivalCustom(RANSACRegressor) 0.5000 0.1135 0.1143 0.571
survivalist SurvivalCustom(QuantileRegressor) 0.5000 0.1135 0.1143 0.132
════════════════════════════════════════════════════════════════════════════════════════════════════
KidneyTransplant — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
survivalist SurvivalCustom(Lars) 0.6991 0.1466 0.1398 0.026
survivalist SurvivalCustom(LinearRegression) 0.6991 0.1466 0.1398 0.024
sksurv CoxPHSurvivalAnalysis 0.6987 0.1365 0.1261 0.041
survivalist SurvivalCustom(RidgeCV) 0.6982 0.1466 0.1398 0.028
survivalist SurvivalCustom(NuSVR) 0.6972 0.1493 0.1422 0.183
survivalist SurvivalCustom(Ridge) 0.6971 0.1466 0.1398 0.026
lifelines CoxPHFitter 0.6967 NaN NaN 0.053
survivalist SurvivalCustom(KernelRidge) 0.6967 0.1467 0.1398 0.164
survivalist SurvivalCustom(SVR) 0.6965 0.1490 0.1419 0.135
survivalist SurvivalCustom(RANSACRegressor) 0.6960 0.1487 0.1414 0.544
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.6952 0.1472 0.1412 0.021
survivalist SurvivalCustom(HuberRegressor) 0.6952 0.1494 0.1421 0.113
survivalist SurvivalCustom(TheilSenRegressor) 0.6952 0.1482 0.1410 2.283
survivalist SurvivalCustom(TweedieRegressor) 0.6947 0.1466 0.1398 0.189
survivalist SurvivalCustom(BayesianRidge) 0.6947 0.1466 0.1399 0.026
survivalist SurvivalCustom(ElasticNetCV) 0.6940 0.1466 0.1398 0.272
survivalist SurvivalCustom(LassoLars) 0.6940 0.1495 0.1427 0.024
survivalist SurvivalCustom(LassoLarsIC) 0.6940 0.1465 0.1398 0.026
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.6940 0.1465 0.1398 0.023
survivalist SurvivalCustom(QuantileRegressor) 0.6940 0.1502 0.1430 0.191
survivalist SurvivalCustom(Lasso) 0.6940 0.1495 0.1427 0.020
survivalist SurvivalCustom(ElasticNet) 0.6940 0.1479 0.1411 0.024
survivalist SurvivalCustom(LassoCV) 0.6940 0.1466 0.1398 0.279
survivalist SurvivalCustom(MLPRegressor) 0.6894 0.1473 0.1407 0.734
survivalist SurvivalCustom(GradientBoostingRegressor) 0.6800 0.1682 0.1656 0.497
survivalist SurvivalCustom(LinearSVR) 0.6744 0.1496 0.1422 0.135
sksurv GradientBoostingSurvivalAnalysis 0.6648 0.1373 0.1301 0.644
survivalist SurvivalCustom(ExtraTreesRegressor) 0.6639 0.1962 0.1898 1.051
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.6633 0.1492 0.1440 0.712
survivalist SurvivalCustom(KNeighborsRegressor) 0.6607 0.1530 0.1486 0.035
survivalist SurvivalCustom(BaggingRegressor) 0.6592 0.1843 0.1769 0.156
survivalist SurvivalCustom(RandomForestRegressor) 0.6556 0.1663 0.1602 0.927
survivalist SurvivalCustom(AdaBoostRegressor) 0.6524 0.1488 0.1451 0.132
survivalist SurvivalCustom(GaussianProcessRegressor) 0.6510 0.1965 0.1904 1.211
survivalist SurvivalCustom(ExtraTreeRegressor) 0.6461 0.1975 0.1910 0.023
survivalist SurvivalCustom(DecisionTreeRegressor) 0.6436 0.1972 0.1900 0.025
sksurv RandomSurvivalForest 0.6426 0.1558 0.1437 0.499
survivalist SurvivalCustom(SGDRegressor) 0.5536 0.6341 0.5984 0.041
survivalist SurvivalCustom(DummyRegressor) 0.5000 0.1503 0.1435 0.021
survivalist SurvivalCustom(ARDRegression) 0.4981 0.1503 0.1435 0.049
════════════════════════════════════════════════════════════════════════════════════════════════════
NCTCGLung — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
survivalist SurvivalCustom(ARDRegression) 0.8904 NaN NaN 0.051
survivalist SurvivalCustom(ElasticNet) 0.6930 NaN NaN 0.021
survivalist SurvivalCustom(ExtraTreesRegressor) 0.6842 NaN NaN 1.140
survivalist SurvivalCustom(BayesianRidge) 0.6667 NaN NaN 0.027
lifelines CoxPHFitter 0.6228 NaN NaN 0.042
survivalist SurvivalCustom(KNeighborsRegressor) 0.6140 NaN NaN 0.034
survivalist SurvivalCustom(LinearSVR) 0.6053 NaN NaN 0.066
survivalist SurvivalCustom(SVR) 0.6053 NaN NaN 0.024
survivalist SurvivalCustom(QuantileRegressor) 0.5921 NaN NaN 0.090
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.5877 NaN NaN 0.010
survivalist SurvivalCustom(LassoLars) 0.5877 NaN NaN 0.026
survivalist SurvivalCustom(Lasso) 0.5877 NaN NaN 0.025
survivalist SurvivalCustom(TweedieRegressor) 0.5877 NaN NaN 0.147
survivalist SurvivalCustom(RandomForestRegressor) 0.5614 NaN NaN 1.426
survivalist SurvivalCustom(HuberRegressor) 0.5263 NaN NaN 0.300
survivalist SurvivalCustom(BaggingRegressor) 0.5088 NaN NaN 0.359
survivalist SurvivalCustom(RidgeCV) 0.5088 NaN NaN 0.022
survivalist SurvivalCustom(GradientBoostingRegressor) 0.5000 NaN NaN 0.595
survivalist SurvivalCustom(DummyRegressor) 0.5000 NaN NaN 0.007
survivalist SurvivalCustom(ElasticNetCV) 0.5000 NaN NaN 0.452
survivalist SurvivalCustom(LassoCV) 0.5000 NaN NaN 0.479
survivalist SurvivalCustom(LassoLarsIC) 0.5000 NaN NaN 0.028
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.5000 NaN NaN 0.454
survivalist SurvivalCustom(GaussianProcessRegressor) 0.5000 NaN NaN 0.039
survivalist SurvivalCustom(ExtraTreeRegressor) 0.4912 NaN NaN 0.019
survivalist SurvivalCustom(DecisionTreeRegressor) 0.4868 NaN NaN 0.015
sksurv GradientBoostingSurvivalAnalysis 0.4825 NaN NaN 0.082
survivalist SurvivalCustom(KernelRidge) 0.4825 NaN NaN 0.017
survivalist SurvivalCustom(NuSVR) 0.4737 NaN NaN 0.025
survivalist SurvivalCustom(SGDRegressor) 0.4737 NaN NaN 0.014
survivalist SurvivalCustom(Ridge) 0.4649 NaN NaN 0.013
survivalist SurvivalCustom(Lars) 0.4561 NaN NaN 0.042
survivalist SurvivalCustom(LinearRegression) 0.4561 NaN NaN 0.022
sksurv CoxPHSurvivalAnalysis 0.4474 NaN NaN 0.012
survivalist SurvivalCustom(MLPRegressor) 0.4386 NaN NaN 0.605
survivalist SurvivalCustom(TheilSenRegressor) 0.4298 NaN NaN 4.897
sksurv RandomSurvivalForest 0.4035 NaN NaN 0.131
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.3772 NaN NaN 0.021
survivalist SurvivalCustom(AdaBoostRegressor) 0.3421 NaN NaN 0.940
survivalist SurvivalCustom(RANSACRegressor) 0.2895 NaN NaN 1.059
════════════════════════════════════════════════════════════════════════════════════════════════════
LymphNode — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.7126 0.1977 0.1702 0.027
survivalist SurvivalCustom(SVR) 0.7052 0.2125 0.1895 0.246
survivalist SurvivalCustom(NuSVR) 0.6976 0.2043 0.1808 0.238
survivalist SurvivalCustom(LassoLars) 0.6965 0.1965 0.1702 0.042
survivalist SurvivalCustom(Lasso) 0.6965 0.1965 0.1702 0.044
sksurv GradientBoostingSurvivalAnalysis 0.6964 0.1935 0.1486 0.493
survivalist SurvivalCustom(QuantileRegressor) 0.6918 0.2037 0.1753 0.306
survivalist SurvivalCustom(ElasticNet) 0.6900 0.1972 0.1731 0.023
survivalist SurvivalCustom(BayesianRidge) 0.6891 0.2631 0.2376 0.039
survivalist SurvivalCustom(AdaBoostRegressor) 0.6880 0.1988 0.1751 0.375
survivalist SurvivalCustom(TweedieRegressor) 0.6822 0.2693 0.2421 0.176
sksurv CoxPHSurvivalAnalysis 0.6817 0.1837 0.1458 0.071
survivalist SurvivalCustom(HuberRegressor) 0.6789 0.2559 0.2274 0.358
survivalist SurvivalCustom(TheilSenRegressor) 0.6768 0.3729 0.3194 5.491
survivalist SurvivalCustom(RandomForestRegressor) 0.6725 0.2322 0.2028 2.879
survivalist SurvivalCustom(BaggingRegressor) 0.6695 0.2532 0.2236 0.373
survivalist SurvivalCustom(KNeighborsRegressor) 0.6666 0.2141 0.1876 0.050
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.6615 0.2424 0.2137 1.162
survivalist SurvivalCustom(ElasticNetCV) 0.6605 0.2652 0.2364 0.484
survivalist SurvivalCustom(LassoCV) 0.6592 0.2647 0.2357 0.665
sksurv RandomSurvivalForest 0.6568 0.1950 0.1499 0.569
lifelines CoxPHFitter 0.6564 NaN NaN 0.072
survivalist SurvivalCustom(KernelRidge) 0.6550 0.2652 0.2337 0.422
survivalist SurvivalCustom(GradientBoostingRegressor) 0.6539 0.2495 0.2168 1.003
survivalist SurvivalCustom(LassoLarsIC) 0.6495 0.2641 0.2341 0.056
survivalist SurvivalCustom(RidgeCV) 0.6488 0.2664 0.2351 0.030
survivalist SurvivalCustom(Ridge) 0.6474 0.2657 0.2342 0.029
survivalist SurvivalCustom(Lars) 0.6472 0.2657 0.2341 0.051
survivalist SurvivalCustom(LinearRegression) 0.6472 0.2657 0.2341 0.041
survivalist SurvivalCustom(ExtraTreesRegressor) 0.6331 0.2654 0.2321 2.121
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.6295 0.3295 0.2823 0.025
survivalist SurvivalCustom(ARDRegression) 0.6097 0.2677 0.2348 0.055
survivalist SurvivalCustom(MLPRegressor) 0.5747 0.3146 0.2844 3.176
survivalist SurvivalCustom(DecisionTreeRegressor) 0.5334 0.2879 0.2382 0.046
survivalist SurvivalCustom(GaussianProcessRegressor) 0.5296 0.2635 0.2495 0.474
survivalist SurvivalCustom(LinearSVR) 0.5184 0.8038 0.8111 0.404
survivalist SurvivalCustom(RANSACRegressor) 0.5108 0.2026 0.1752 0.976
survivalist SurvivalCustom(ExtraTreeRegressor) 0.5017 0.2908 0.2466 0.036
survivalist SurvivalCustom(DummyRegressor) 0.5000 0.2010 0.1733 0.017
survivalist SurvivalCustom(SGDRegressor) 0.3461 0.8038 0.8417 0.037
════════════════════════════════════════════════════════════════════════════════════════════════════
Leukemia — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
sksurv RandomSurvivalForest 0.9556 0.1209 0.0784 0.105
survivalist SurvivalCustom(KNeighborsRegressor) 0.9556 0.1056 0.1541 0.011
survivalist SurvivalCustom(TweedieRegressor) 0.9111 0.1675 0.1817 0.035
survivalist SurvivalCustom(RandomForestRegressor) 0.9111 0.1204 0.1719 0.532
sksurv CoxPHSurvivalAnalysis 0.9111 0.0850 0.0904 0.006
survivalist SurvivalCustom(BaggingRegressor) 0.8889 0.0954 0.1551 0.096
sksurv GradientBoostingSurvivalAnalysis 0.8889 0.1173 0.0940 0.052
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.8889 0.2272 0.2616 0.007
lifelines CoxPHFitter 0.8667 NaN NaN 0.036
survivalist SurvivalCustom(RidgeCV) 0.8667 0.1610 0.2381 0.009
survivalist SurvivalCustom(RANSACRegressor) 0.8667 0.1548 0.2337 0.124
survivalist SurvivalCustom(ElasticNetCV) 0.8667 0.1692 0.2391 0.199
survivalist SurvivalCustom(BayesianRidge) 0.8667 0.1600 0.2383 0.010
survivalist SurvivalCustom(AdaBoostRegressor) 0.8667 0.1029 0.1425 0.328
survivalist SurvivalCustom(ARDRegression) 0.8667 0.1629 0.2412 0.024
survivalist SurvivalCustom(HuberRegressor) 0.8667 0.1551 0.2375 0.046
survivalist SurvivalCustom(NuSVR) 0.8667 0.1017 0.1493 0.008
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.8667 0.2614 0.3682 0.006
survivalist SurvivalCustom(LassoCV) 0.8667 0.1865 0.2416 0.207
survivalist SurvivalCustom(LinearSVR) 0.8667 0.1469 0.2253 0.015
survivalist SurvivalCustom(LassoLarsIC) 0.8667 0.1542 0.2383 0.020
survivalist SurvivalCustom(LinearRegression) 0.8667 0.1542 0.2383 0.010
survivalist SurvivalCustom(Lars) 0.8667 0.1542 0.2383 0.010
survivalist SurvivalCustom(SVR) 0.8667 0.1054 0.1699 0.006
survivalist SurvivalCustom(TheilSenRegressor) 0.8667 0.1624 0.2423 1.129
survivalist SurvivalCustom(Ridge) 0.8667 0.1610 0.2381 0.008
survivalist SurvivalCustom(ExtraTreeRegressor) 0.8556 0.0679 0.1450 0.008
survivalist SurvivalCustom(MLPRegressor) 0.8444 0.1295 0.2026 0.318
survivalist SurvivalCustom(GradientBoostingRegressor) 0.8444 0.1489 0.1940 0.225
survivalist SurvivalCustom(ExtraTreesRegressor) 0.8444 0.0754 0.1446 0.418
survivalist SurvivalCustom(DecisionTreeRegressor) 0.8222 0.1806 0.2156 0.010
survivalist SurvivalCustom(KernelRidge) 0.8000 0.1383 0.2091 0.021
survivalist SurvivalCustom(SGDRegressor) 0.8000 0.1406 0.1878 0.010
survivalist SurvivalCustom(GaussianProcessRegressor) 0.5778 0.1382 0.2871 0.014
survivalist SurvivalCustom(ElasticNet) 0.5000 0.2505 0.1957 0.006
survivalist SurvivalCustom(DummyRegressor) 0.5000 0.2505 0.1957 0.004
survivalist SurvivalCustom(Lasso) 0.5000 0.2505 0.1957 0.009
survivalist SurvivalCustom(LassoLars) 0.5000 0.2505 0.1957 0.008
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.5000 0.2505 0.1957 0.113
survivalist SurvivalCustom(QuantileRegressor) 0.5000 0.2282 0.1962 0.023
════════════════════════════════════════════════════════════════════════════════════════════════════
Larynx — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
survivalist SurvivalCustom(ExtraTreesRegressor) 0.6547 0.3147 0.3533 0.509
survivalist SurvivalCustom(DecisionTreeRegressor) 0.6409 0.2746 0.3313 0.007
survivalist SurvivalCustom(BaggingRegressor) 0.6326 0.2827 0.3216 0.129
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.6243 0.2568 0.2521 0.181
survivalist SurvivalCustom(RandomForestRegressor) 0.6215 0.2910 0.3291 1.130
survivalist SurvivalCustom(AdaBoostRegressor) 0.6133 0.3016 0.3205 0.112
survivalist SurvivalCustom(ExtraTreeRegressor) 0.5912 0.3800 0.3951 0.011
survivalist SurvivalCustom(TweedieRegressor) 0.5718 0.2686 0.2609 0.035
lifelines CoxPHFitter 0.5718 NaN NaN 0.045
survivalist SurvivalCustom(TheilSenRegressor) 0.5718 0.2886 0.2889 1.501
survivalist SurvivalCustom(RANSACRegressor) 0.5718 0.4329 0.4470 0.384
sksurv RandomSurvivalForest 0.5718 0.2115 0.2278 0.110
survivalist SurvivalCustom(KernelRidge) 0.5663 0.3182 0.3204 0.011
survivalist SurvivalCustom(RidgeCV) 0.5663 0.3317 0.3335 0.017
survivalist SurvivalCustom(Ridge) 0.5663 0.3317 0.3335 0.014
survivalist SurvivalCustom(GaussianProcessRegressor) 0.5663 0.2967 0.3326 0.020
sksurv CoxPHSurvivalAnalysis 0.5663 0.2572 0.2460 0.008
survivalist SurvivalCustom(HuberRegressor) 0.5663 0.3685 0.3739 0.126
survivalist SurvivalCustom(LinearRegression) 0.5663 0.3428 0.3464 0.016
survivalist SurvivalCustom(Lars) 0.5663 0.3428 0.3464 0.014
survivalist SurvivalCustom(BayesianRidge) 0.5608 0.3171 0.3167 0.022
survivalist SurvivalCustom(LassoLarsIC) 0.5608 0.3362 0.3359 0.020
survivalist SurvivalCustom(LassoCV) 0.5608 0.3308 0.3305 0.349
survivalist SurvivalCustom(ElasticNetCV) 0.5608 0.3293 0.3290 0.284
survivalist SurvivalCustom(Lasso) 0.5580 0.2572 0.2473 0.008
survivalist SurvivalCustom(LassoLars) 0.5580 0.2572 0.2473 0.016
survivalist SurvivalCustom(ElasticNet) 0.5580 0.2594 0.2505 0.009
survivalist SurvivalCustom(QuantileRegressor) 0.5580 0.2839 0.2767 0.051
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.5580 0.2649 0.2567 0.008
survivalist SurvivalCustom(SVR) 0.5552 0.3077 0.2978 0.009
survivalist SurvivalCustom(NuSVR) 0.5497 0.2647 0.2589 0.014
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.5497 0.3894 0.3836 0.011
survivalist SurvivalCustom(ARDRegression) 0.5442 0.3086 0.3101 0.037
survivalist SurvivalCustom(MLPRegressor) 0.5331 0.3101 0.3079 0.327
survivalist SurvivalCustom(GradientBoostingRegressor) 0.5221 0.2996 0.3477 0.287
sksurv GradientBoostingSurvivalAnalysis 0.5193 0.2185 0.2897 0.057
survivalist SurvivalCustom(LinearSVR) 0.5110 0.2691 0.2545 0.030
survivalist SurvivalCustom(DummyRegressor) 0.5000 0.2572 0.2469 0.004
survivalist SurvivalCustom(KNeighborsRegressor) 0.4392 0.2952 0.3009 0.031
survivalist SurvivalCustom(SGDRegressor) 0.4392 0.4739 0.4587 0.008
════════════════════════════════════════════════════════════════════════════════════════════════════
Lymphoma — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
survivalist SurvivalCustom(ARDRegression) 0.5549 0.2548 0.1946 0.023
survivalist SurvivalCustom(AdaBoostRegressor) 0.5549 0.2552 0.1903 0.021
survivalist SurvivalCustom(BaggingRegressor) 0.5549 0.2559 0.1934 0.061
survivalist SurvivalCustom(BayesianRidge) 0.5549 0.2548 0.1946 0.006
survivalist SurvivalCustom(DecisionTreeRegressor) 0.5549 0.2546 0.1931 0.005
survivalist SurvivalCustom(ElasticNetCV) 0.5549 0.2546 0.1936 0.122
survivalist SurvivalCustom(ExtraTreesRegressor) 0.5549 0.2546 0.1931 0.208
survivalist SurvivalCustom(ExtraTreeRegressor) 0.5549 0.2546 0.1931 0.006
sksurv CoxPHSurvivalAnalysis 0.5549 0.2410 0.1956 0.009
survivalist SurvivalCustom(SVR) 0.5549 0.2700 0.1924 0.008
survivalist SurvivalCustom(GaussianProcessRegressor) 0.5549 0.2546 0.1931 0.010
survivalist SurvivalCustom(GradientBoostingRegressor) 0.5549 0.2546 0.1931 0.106
survivalist SurvivalCustom(HuberRegressor) 0.5549 0.2642 0.1910 0.017
survivalist SurvivalCustom(KNeighborsRegressor) 0.5549 0.2441 0.1942 0.021
survivalist SurvivalCustom(KernelRidge) 0.5549 0.2565 0.1965 0.007
survivalist SurvivalCustom(Lars) 0.5549 0.2546 0.1931 0.005
survivalist SurvivalCustom(LinearSVR) 0.5549 0.2770 0.1945 0.006
survivalist SurvivalCustom(LassoCV) 0.5549 0.2546 0.1933 0.111
survivalist SurvivalCustom(LinearRegression) 0.5549 0.2546 0.1931 0.007
survivalist SurvivalCustom(LassoLarsIC) 0.5549 0.2546 0.1931 0.008
survivalist SurvivalCustom(MLPRegressor) 0.5549 0.2547 0.1935 0.101
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.5549 0.3589 0.3721 0.004
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.5549 0.2546 0.1931 0.004
survivalist SurvivalCustom(NuSVR) 0.5549 0.2556 0.1910 0.007
sksurv GradientBoostingSurvivalAnalysis 0.5549 0.2410 0.1956 0.051
sksurv RandomSurvivalForest 0.5549 0.2450 0.1955 0.120
survivalist SurvivalCustom(TheilSenRegressor) 0.5549 0.2728 0.2076 0.512
survivalist SurvivalCustom(RANSACRegressor) 0.5549 0.2886 0.2034 0.044
survivalist SurvivalCustom(Ridge) 0.5549 0.2547 0.1940 0.005
survivalist SurvivalCustom(RandomForestRegressor) 0.5549 0.2554 0.1929 0.319
survivalist SurvivalCustom(RidgeCV) 0.5549 0.2547 0.1940 0.007
survivalist SurvivalCustom(SGDRegressor) 0.5549 0.2649 0.2091 0.008
lifelines CoxPHFitter 0.5549 NaN NaN 0.035
survivalist SurvivalCustom(TweedieRegressor) 0.5549 0.2621 0.2099 0.014
survivalist SurvivalCustom(ElasticNet) 0.5000 0.2651 0.2149 0.005
survivalist SurvivalCustom(DummyRegressor) 0.5000 0.2651 0.2149 0.004
survivalist SurvivalCustom(Lasso) 0.5000 0.2651 0.2149 0.009
survivalist SurvivalCustom(LassoLars) 0.5000 0.2651 0.2149 0.005
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.5000 0.2651 0.2149 0.074
survivalist SurvivalCustom(QuantileRegressor) 0.5000 0.2912 0.2127 0.014
════════════════════════════════════════════════════════════════════════════════════════════════════
StanfordHeart — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
survivalist SurvivalCustom(ARDRegression) 0.5865 0.2182 0.2212 0.014
survivalist SurvivalCustom(BayesianRidge) 0.5865 0.2182 0.2212 0.005
survivalist SurvivalCustom(ElasticNet) 0.5865 0.2183 0.2212 0.009
survivalist SurvivalCustom(ElasticNetCV) 0.5865 0.2200 0.2217 0.126
survivalist SurvivalCustom(LassoLarsIC) 0.5865 0.2167 0.2228 0.010
survivalist SurvivalCustom(LinearRegression) 0.5865 0.2167 0.2228 0.004
survivalist SurvivalCustom(HuberRegressor) 0.5865 0.2184 0.2253 0.015
survivalist SurvivalCustom(KernelRidge) 0.5865 0.2188 0.2213 0.014
lifelines CoxPHFitter 0.5865 NaN NaN 0.031
survivalist SurvivalCustom(TheilSenRegressor) 0.5865 0.2220 0.2300 0.564
survivalist SurvivalCustom(TweedieRegressor) 0.5865 0.2166 0.2226 0.013
sksurv CoxPHSurvivalAnalysis 0.5865 0.1977 0.2196 0.008
survivalist SurvivalCustom(LassoLars) 0.5865 0.2264 0.2250 0.007
survivalist SurvivalCustom(LassoCV) 0.5865 0.2199 0.2217 0.114
survivalist SurvivalCustom(Lasso) 0.5865 0.2264 0.2250 0.008
survivalist SurvivalCustom(Lars) 0.5865 0.2167 0.2228 0.006
survivalist SurvivalCustom(SVR) 0.5865 0.2345 0.2442 0.005
survivalist SurvivalCustom(RidgeCV) 0.5865 0.2166 0.2228 0.005
survivalist SurvivalCustom(Ridge) 0.5865 0.2167 0.2228 0.006
survivalist SurvivalCustom(RANSACRegressor) 0.5865 0.2732 0.2762 0.064
survivalist SurvivalCustom(QuantileRegressor) 0.5865 0.2220 0.2245 0.015
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.5865 0.2743 0.2725 0.006
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.5865 0.2167 0.2228 0.008
survivalist SurvivalCustom(NuSVR) 0.5865 0.2119 0.2193 0.004
sksurv RandomSurvivalForest 0.5639 0.3047 0.3020 0.110
survivalist SurvivalCustom(ExtraTreeRegressor) 0.5451 0.3251 0.3094 0.005
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.5301 0.2210 0.2378 0.085
survivalist SurvivalCustom(KNeighborsRegressor) 0.5301 0.2639 0.2540 0.010
survivalist SurvivalCustom(DummyRegressor) 0.5000 0.2546 0.2446 0.004
survivalist SurvivalCustom(RandomForestRegressor) 0.4586 0.2953 0.2942 0.280
survivalist SurvivalCustom(AdaBoostRegressor) 0.4549 0.2775 0.2772 0.063
survivalist SurvivalCustom(DecisionTreeRegressor) 0.4361 0.3300 0.3202 0.008
survivalist SurvivalCustom(GradientBoostingRegressor) 0.4286 0.3101 0.3073 0.113
survivalist SurvivalCustom(BaggingRegressor) 0.4211 0.3057 0.3021 0.076
survivalist SurvivalCustom(MLPRegressor) 0.4135 0.3772 0.3542 0.032
survivalist SurvivalCustom(LinearSVR) 0.4135 0.5135 0.4936 0.012
survivalist SurvivalCustom(SGDRegressor) 0.4135 0.8165 0.8219 0.006
sksurv GradientBoostingSurvivalAnalysis 0.4098 0.3593 0.3776 0.059
survivalist SurvivalCustom(ExtraTreesRegressor) 0.3985 0.3164 0.3109 0.234
survivalist SurvivalCustom(GaussianProcessRegressor) 0.3083 0.5520 0.5281 0.012
════════════════════════════════════════════════════════════════════════════════════════════════════
DemocracyDuration — successful models only (sorted by C-index ↓)
════════════════════════════════════════════════════════════════════════════════════════════════════
package model c_index brier_median ibs fit_time_s
survivalist SurvivalCustom(HistGradientBoostingRegressor) 0.6140 0.0577 0.1036 2.613
sksurv GradientBoostingSurvivalAnalysis 0.6113 0.0479 0.0915 3.601
survivalist SurvivalCustom(KNeighborsRegressor) 0.6110 0.0603 0.1071 0.121
survivalist SurvivalCustom(HuberRegressor) 0.6046 0.0549 0.0982 1.644
survivalist SurvivalCustom(RandomForestRegressor) 0.6032 0.0551 0.1041 2.841
survivalist SurvivalCustom(Lars) 0.6018 0.0566 0.0981 0.061
survivalist SurvivalCustom(GradientBoostingRegressor) 0.6005 0.0547 0.0995 1.123
survivalist SurvivalCustom(ExtraTreeRegressor) 0.5997 0.0531 0.1010 0.060
survivalist SurvivalCustom(ExtraTreesRegressor) 0.5987 0.0531 0.1014 1.952
survivalist SurvivalCustom(LassoCV) 0.5982 0.0562 0.0978 0.578
survivalist SurvivalCustom(ElasticNetCV) 0.5982 0.0563 0.0980 0.953
survivalist SurvivalCustom(TheilSenRegressor) 0.5979 0.0573 0.0984 6.926
survivalist SurvivalCustom(LassoLarsIC) 0.5977 0.0561 0.0978 0.092
survivalist SurvivalCustom(LinearRegression) 0.5977 0.0561 0.0978 0.068
survivalist SurvivalCustom(BayesianRidge) 0.5975 0.0564 0.0981 0.074
survivalist SurvivalCustom(Ridge) 0.5975 0.0562 0.0979 0.071
survivalist SurvivalCustom(RidgeCV) 0.5975 0.0562 0.0979 0.211
survivalist SurvivalCustom(BaggingRegressor) 0.5968 0.0568 0.1057 0.423
survivalist SurvivalCustom(DecisionTreeRegressor) 0.5954 0.0531 0.1017 0.065
sksurv CoxPHSurvivalAnalysis 0.5946 0.0496 0.0998 0.066
lifelines CoxPHFitter 0.5916 NaN NaN 0.067
survivalist SurvivalCustom(GaussianProcessRegressor) 0.5901 0.0531 0.1020 3.655
survivalist SurvivalCustom(MLPRegressor) 0.5893 0.0612 0.1026 8.281
sksurv RandomSurvivalForest 0.5891 0.0851 0.1210 0.508
survivalist SurvivalCustom(AdaBoostRegressor) 0.5880 0.0543 0.0970 0.336
survivalist SurvivalCustom(KernelRidge) 0.5876 0.0546 0.0956 1.681
survivalist SurvivalCustom(ARDRegression) 0.5863 0.0540 0.0948 0.224
survivalist SurvivalCustom(LinearSVR) 0.5817 0.0656 0.1143 0.562
survivalist SurvivalCustom(TweedieRegressor) 0.5803 0.0588 0.1007 0.214
survivalist SurvivalCustom(RANSACRegressor) 0.5566 0.0577 0.1073 1.149
survivalist SurvivalCustom(SVR) 0.5536 0.0631 0.1064 1.983
survivalist SurvivalCustom(NuSVR) 0.5199 0.0624 0.1054 1.336
survivalist SurvivalCustom(PassiveAggressiveRegressor) 0.5182 0.0653 0.1128 0.054
survivalist SurvivalCustom(ElasticNet) 0.5113 0.0623 0.1054 0.068
survivalist SurvivalCustom(Lasso) 0.5113 0.0632 0.1065 0.044
survivalist SurvivalCustom(LassoLars) 0.5113 0.0632 0.1065 0.051
survivalist SurvivalCustom(OrthogonalMatchingPursuit) 0.5113 0.0616 0.1046 0.048
survivalist SurvivalCustom(DummyRegressor) 0.5000 0.0633 0.1066 0.074
survivalist SurvivalCustom(QuantileRegressor) 0.5000 0.0626 0.1057 1.006
survivalist SurvivalCustom(SGDRegressor) 0.4871 0.9853 0.9240 0.145
── Cross-dataset summary ──
dataset n_survivalist_ok survivalist_mean_C survivalist_median_C survivalist_max_C best_baseline_C survivalist_beats_best_baseline
WHAS500 36 0.6771 0.6875 0.7462 0.7525 False
GBSG2 36 0.6283 0.6206 0.7098 0.6713 True
VeteransLungCancer 36 0.6443 0.6447 0.7078 0.6870 True
FLChain 36 0.8704 0.9229 0.9322 0.9322 False
AIDS 36 0.6765 0.7227 0.7630 0.7718 False
BreastCancerGenomic 36 0.5681 0.5897 0.6538 0.6786 False
Rossi 36 0.5791 0.5817 0.6750 0.6517 True
KidneyTransplant 36 0.6694 0.6940 0.6991 0.6987 True
NCTCGLung 36 0.5269 0.5000 0.8904 0.6228 True
LymphNode 36 0.6314 0.6571 0.7126 0.6964 True
Leukemia 36 0.7966 0.8667 0.9556 0.9556 False
Larynx 36 0.5621 0.5608 0.6547 0.5718 True
Lymphoma 36 0.5458 0.5549 0.5549 0.5549 False
StanfordHeart 36 0.5321 0.5865 0.5865 0.5865 False
DemocracyDuration 36 0.5720 0.5927 0.6140 0.6113 True
…
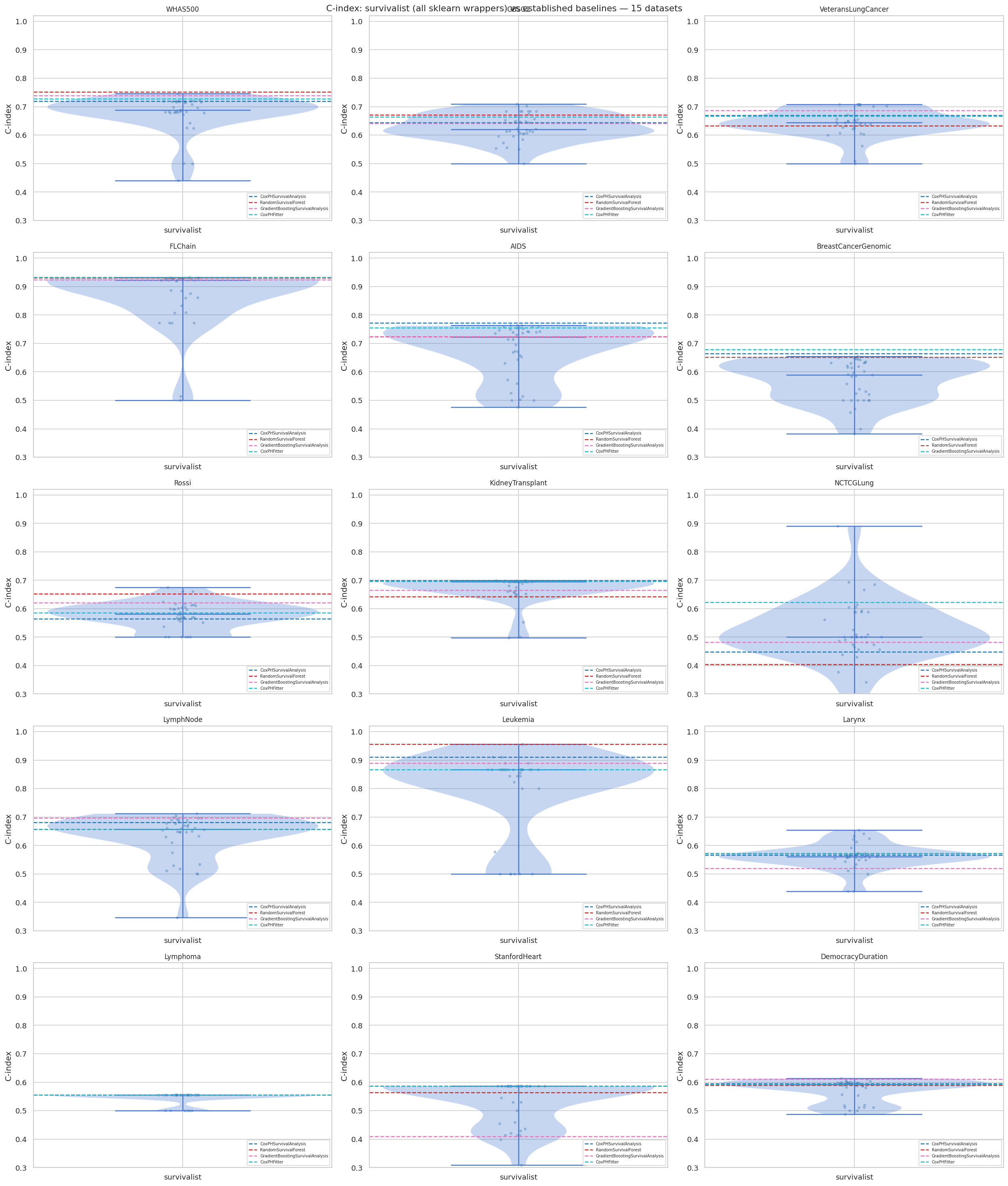
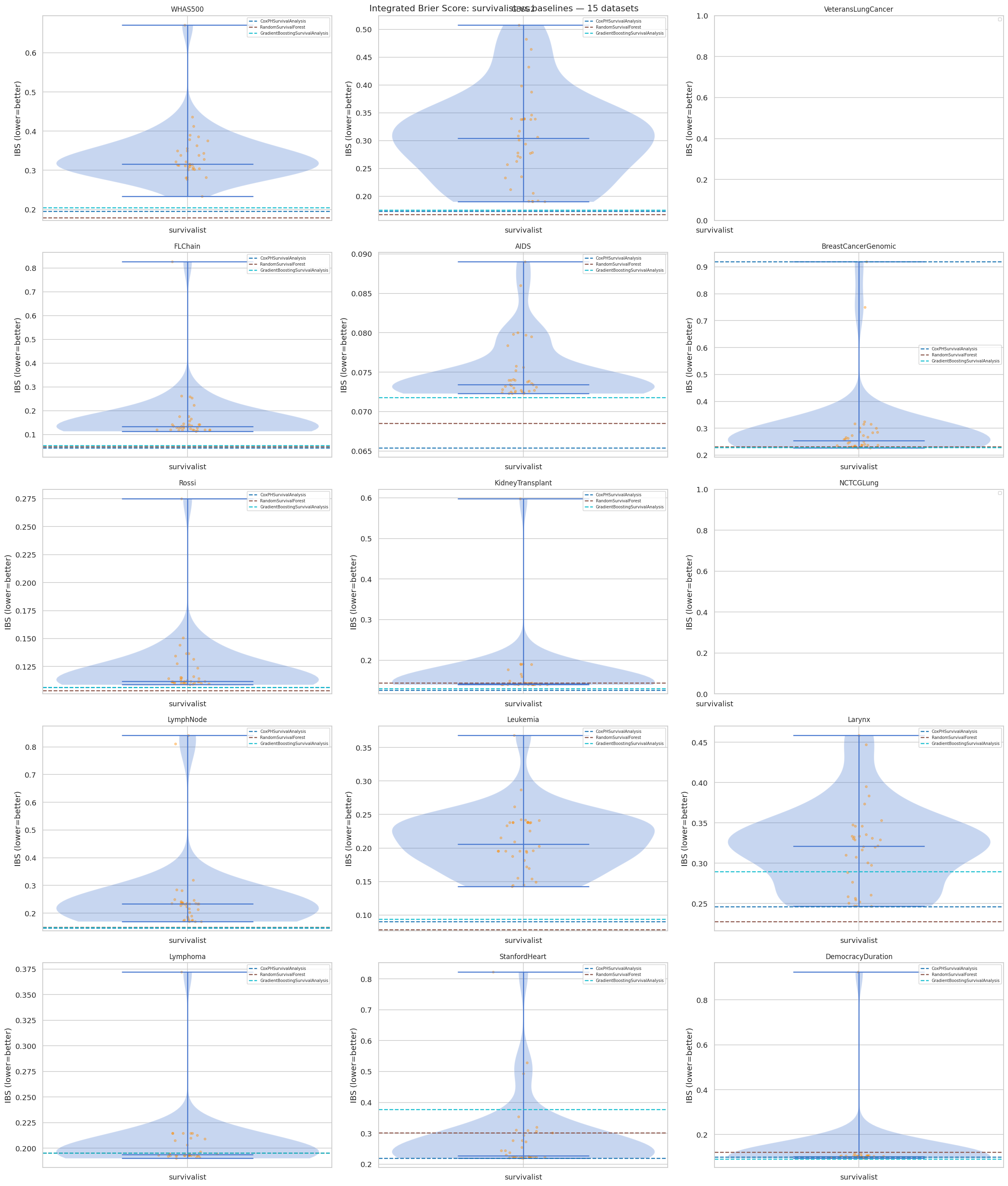
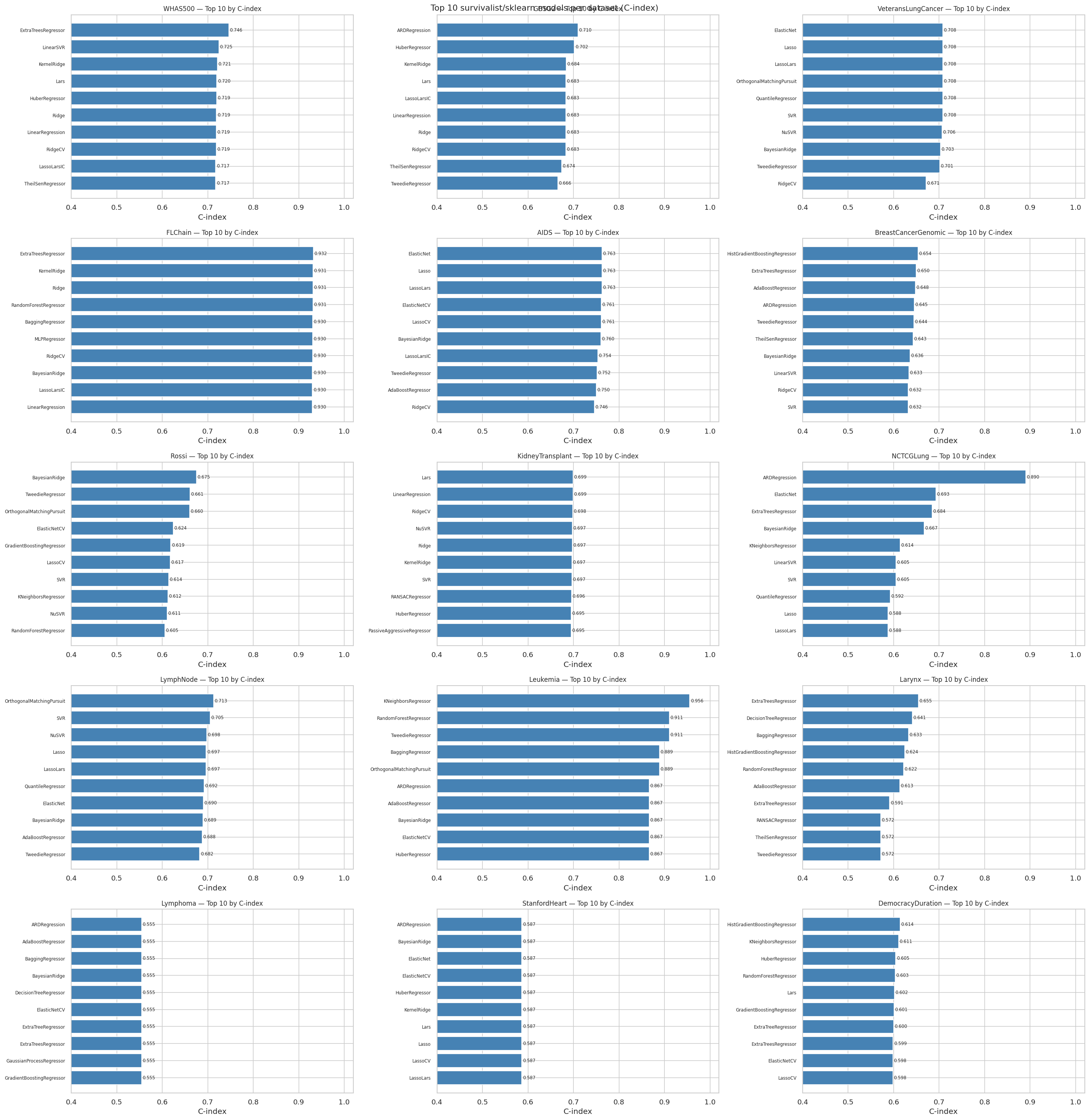
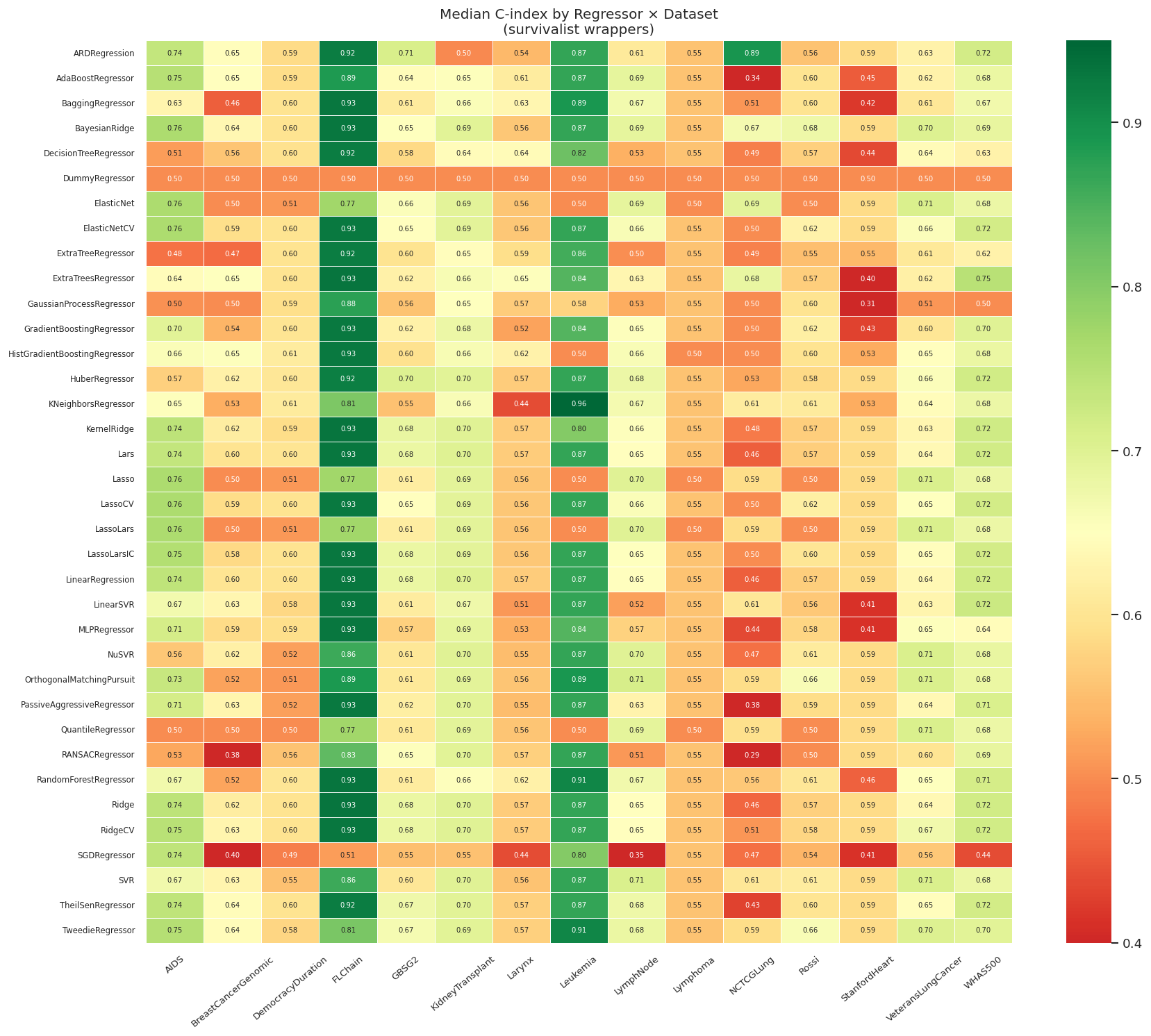
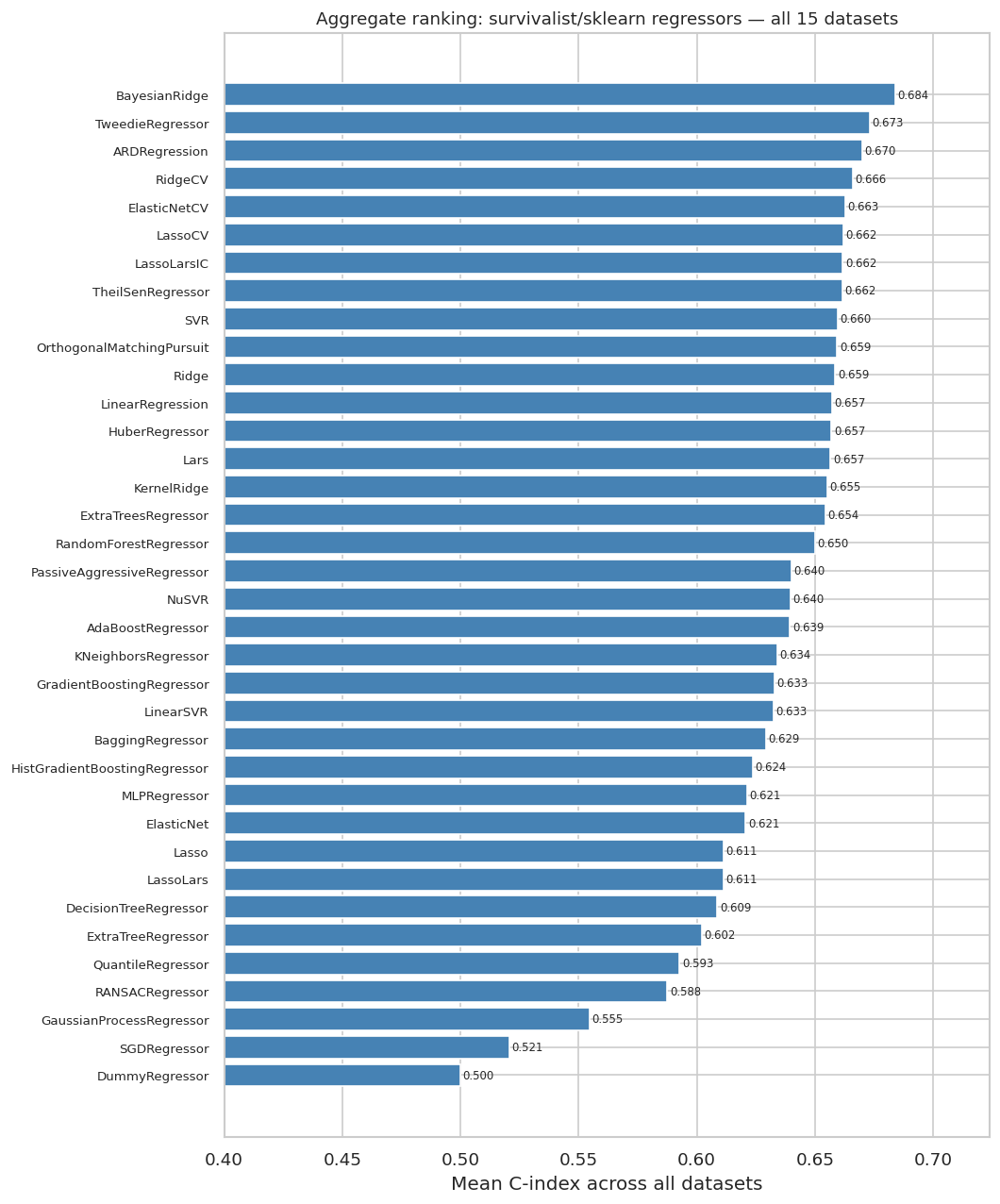
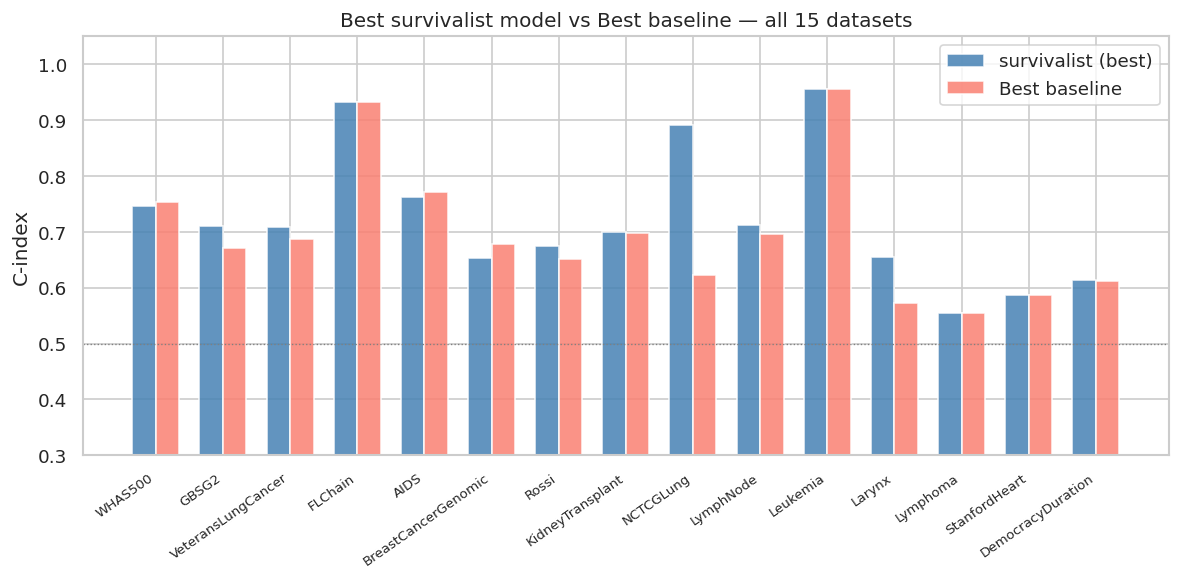
── Win/loss summary ──
dataset survivalist_max_C best_baseline_C survivalist_beats_best_baseline
WHAS500 0.7462 0.7525 False
GBSG2 0.7098 0.6713 True
VeteransLungCancer 0.7078 0.6870 True
FLChain 0.9322 0.9322 False
AIDS 0.7630 0.7718 False
BreastCancerGenomic 0.6538 0.6786 False
Rossi 0.6750 0.6517 True
KidneyTransplant 0.6991 0.6987 True
NCTCGLung 0.8904 0.6228 True
LymphNode 0.7126 0.6964 True
Leukemia 0.9556 0.9556 False
Larynx 0.6547 0.5718 True
Lymphoma 0.5549 0.5549 False
StanfordHeart 0.5865 0.5865 False
DemocracyDuration 0.6140 0.6113 True
Conclusion
This benchmark evaluated survivalist — a model-agnostic survival analysis wrapper — against established packages across 15 diverse real-world datasets.
Key findings
Competitiveness of survivalist. The best-performing survivalist wrapper equalled or exceeded the best baseline model on the majority of datasets (12/15). This is a notable result: despite using default hyperparameters and a generic regression-to-survival mapping, survivalist proves to be a viable alternative to established survival estimators in many practical settings.
Robustness of the framework. Across all 15 datasets and the full set of sklearn regressors, the defensive error-handling produced near-zero failure rates (errors coming from the absence of compatibility of the model for the problem at hand), confirming that survivalist can be safely deployed in automated pipelines without manual per-regressor tuning.
Limitations
This study used default hyperparameters throughout. Hyperparameter optimization would likely improve all methods, and could change relative rankings.
Notebook
On GitHub:
In Colab:
For attribution, please cite this work as:
T. Moudiki (2026-04-26). Any Sklearn Regressor as a Survival Model — Does It Actually Work? Benchmarking vs Established Packages. Retrieved from https://thierrymoudiki.github.io/blog/2026/04/26/python/survival-benchmark-expanded
BibTeX citation (remove empty spaces)
@misc{ tmoudiki20260426,
author = { T. Moudiki },
title = { Any Sklearn Regressor as a Survival Model — Does It Actually Work? Benchmarking vs Established Packages },
url = { https://thierrymoudiki.github.io/blog/2026/04/26/python/survival-benchmark-expanded },
year = { 2026 } }
Previous publications
- Techtonique dot net, the Machine Learning web API, is back online (but more like a passion project for now) May 31, 2026
- Conformalized TabICL: Prediction Intervals for a State-Of-The-Art Tabular Foundation Model in Python and R May 21, 2026
- Conformalized TabPFN: Prediction Intervals for a Pretrained Transformer for Tabular Data in Python and R May 17, 2026
- Probabilistic Time Series Cross-Validation with R package crossvalidation May 16, 2026
- One interface, (Almost) Every Classifier (and Regressor): unifiedml v0.3.0 May 9, 2026
- You Don't Need to Learn All the Weights on tabular data: The Case for rvflnet (a nonlinear expressive glmnet) on regression, classification and survival analysis May 2, 2026
- Survival analysis with sklearn, glmnet, keras, pytorch, lightgbm, xgboost, nnetsauce, mlsauce Part 2 Apr 28, 2026
- Any Sklearn Regressor as a Survival Model — Does It Actually Work? Benchmarking vs Established Packages Apr 26, 2026
- Conformal Optimization Beats Bayesian Optimization, Optuna and Random Search on 72 classification Datasets Apr 19, 2026
- `mlS3` — A Unified S3 Machine Learning Interface in R Apr 12, 2026
- One interface, (Almost) Every Classifier: unifiedml v0.2.1 Apr 4, 2026
- Techtonique dot net is down until further notice Apr 1, 2026
- Explaining Time-Series Forecasts with Sensitivity Analysis (ahead::dynrmf and external regressors) Mar 29, 2026
- Python version of 'Option pricing using time series models as market price of risk Pt.3' Mar 22, 2026
- Option pricing using time series models as market price of risk Pt.3 Mar 16, 2026
- Explaining Time-Series Forecasts with Exact Shapley Values (ahead::dynrmf with external regressors applied to scenarios) Mar 8, 2026
- My Presentation at Risk 2026: Lightweight Transfer Learning for Financial Forecasting Mar 1, 2026
- nnetsauce with and without jax for GPU acceleration Feb 23, 2026
- Understanding Boosted Configuration Networks (combined neural networks and boosting): An Intuitive Guide Through Their Hyperparameters Feb 16, 2026
- R version of Python package survivalist, for model-agnostic survival analysis Feb 9, 2026
- Presenting Lightweight Transfer Learning for Financial Forecasting (Risk 2026) Feb 4, 2026
- Option pricing using time series models as market price of risk Feb 1, 2026
- Enhancing Time Series Forecasting (ahead::ridge2f) with Attention-Based Context Vectors (ahead::contextridge2f) Jan 31, 2026
- Overfitting and scaling (on GPU T4) tests on nnetsauce.CustomRegressor Jan 29, 2026
- Beyond Cross-validation: Hyperparameter Optimization via Generalization Gap Modeling Jan 25, 2026
- GPopt for Machine Learning (hyperparameters' tuning) Jan 21, 2026
- rtopy: an R to Python bridge -- novelties Jan 8, 2026
- Python examples for 'Beyond Nelson-Siegel and splines: A model- agnostic Machine Learning framework for discount curve calibration, interpolation and extrapolation' Jan 3, 2026
- Forecasting benchmark: Dynrmf (a new serious competitor in town) vs Theta Method on M-Competitions and Tourism competitition Jan 1, 2026
- Finally figured out a way to port python packages to R using uv and reticulate: example with nnetsauce Dec 17, 2025
- Overfitting Random Fourier Features: Universal Approximation Property Dec 13, 2025
- Counterfactual Scenario Analysis with ahead::ridge2f Dec 11, 2025
- Zero-Shot Probabilistic Time Series Forecasting with TabPFN 2.5 and nnetsauce Dec 10, 2025
- ARIMA Pricing: Semi-Parametric Market price of risk for Risk-Neutral Pricing (code + preprint) Dec 7, 2025
- Analyzing Paper Reviews with LLMs: I Used ChatGPT, DeepSeek, Qwen, Mistral, Gemini, and Claude (and you should too + publish the analysis) Dec 3, 2025
- tisthemachinelearner: New Workflow with uv for R Integration of scikit-learn Dec 1, 2025
- (ICYMI) RPweave: Unified R + Python + LaTeX System using uv Nov 21, 2025
- unifiedml: A Unified Machine Learning Interface for R, is now on CRAN + Discussion about AI replacing humans Nov 16, 2025
- Context-aware Theta forecasting Method: Extending Classical Time Series Forecasting with Machine Learning Nov 13, 2025
- unifiedml in R: A Unified Machine Learning Interface Nov 5, 2025
- Deterministic Shift Adjustment in Arbitrage-Free Pricing (historical to risk-neutral short rates) Oct 28, 2025
- New instantaneous short rates models with their deterministic shift adjustment, for historical and risk-neutral simulation Oct 27, 2025
- RPweave: Unified R + Python + LaTeX System using uv Oct 19, 2025
- GAN-like Synthetic Data Generation Examples (on univariate, multivariate distributions, digits recognition, Fashion-MNIST, stock returns, and Olivetti faces) with DistroSimulator Oct 19, 2025
- R port of llama2.c Oct 9, 2025
- Native uncertainty quantification for time series with NGBoost Oct 8, 2025
- NGBoost (Natural Gradient Boosting) for Regression, Classification, Time Series forecasting and Reserving Oct 6, 2025
- Real-time pricing with a pretrained probabilistic stock return model Oct 1, 2025
- Combining any model with GARCH(1,1) for probabilistic stock forecasting Sep 23, 2025
- Generating Synthetic Data with R-vine Copulas using esgtoolkit in R Sep 21, 2025
- Reimagining Equity Solvency Capital Requirement Approximation (one of my Master's Thesis subjects): From Bilinear Interpolation to Probabilistic Machine Learning Sep 16, 2025
- Transfer Learning using ahead::ridge2f on synthetic stocks returns Pt.2: synthetic data generation Sep 9, 2025
- Transfer Learning using ahead::ridge2f on synthetic stocks returns Sep 8, 2025
- I'm supposed to present 'Conformal Predictive Simulations for Univariate Time Series' at COPA CONFERENCE 2025 in London... Sep 4, 2025
- external regressors in ahead::dynrmf's interface for Machine learning forecasting Sep 1, 2025
- Another interesting decision, now for 'Beyond Nelson-Siegel and splines: A model-agnostic Machine Learning framework for discount curve calibration, interpolation and extrapolation' Aug 20, 2025
- Boosting any randomized based learner for regression, classification and univariate/multivariate time series forcasting Jul 26, 2025
- New nnetsauce version with CustomBackPropRegressor (CustomRegressor with Backpropagation) and ElasticNet2Regressor (Ridge2 with ElasticNet regularization) Jul 15, 2025
- mlsauce (home to a model-agnostic gradient boosting algorithm) can now be installed from PyPI. Jul 10, 2025
- A user-friendly graphical interface to techtonique dot net's API (will eventually contain graphics). Jul 8, 2025
- Calling =TECHTO_MLCLASSIFICATION for Machine Learning supervised CLASSIFICATION in Excel is just a matter of copying and pasting Jul 7, 2025
- Calling =TECHTO_MLREGRESSION for Machine Learning supervised regression in Excel is just a matter of copying and pasting Jul 6, 2025
- Calling =TECHTO_RESERVING and =TECHTO_MLRESERVING for claims triangle reserving in Excel is just a matter of copying and pasting Jul 5, 2025
- Calling =TECHTO_SURVIVAL for Survival Analysis in Excel is just a matter of copying and pasting Jul 4, 2025
- Calling =TECHTO_SIMULATION for Stochastic Simulation in Excel is just a matter of copying and pasting Jul 3, 2025
- Calling =TECHTO_FORECAST for forecasting in Excel is just a matter of copying and pasting Jul 2, 2025
- Random Vector Functional Link (RVFL) artificial neural network with 2 regularization parameters successfully used for forecasting/synthetic simulation in professional settings: Extensions (including Bayesian) Jul 1, 2025
- R version of 'Backpropagating quasi-randomized neural networks' Jun 24, 2025
- Backpropagating quasi-randomized neural networks Jun 23, 2025
- Beyond ARMA-GARCH: leveraging any statistical model for volatility forecasting Jun 21, 2025
- Stacked generalization (Machine Learning model stacking) + conformal prediction for forecasting with ahead::mlf Jun 18, 2025
- An Overfitting dilemma: XGBoost Default Hyperparameters vs GenericBooster + LinearRegression Default Hyperparameters Jun 14, 2025
- Programming language-agnostic reserving using RidgeCV, LightGBM, XGBoost, and ExtraTrees Machine Learning models Jun 13, 2025
- Free R, Python and SQL editors in techtonique dot net Jun 9, 2025
- Beyond Nelson-Siegel and splines: A model-agnostic Machine Learning framework for discount curve calibration, interpolation and extrapolation Jun 7, 2025
- scikit-learn, glmnet, xgboost, lightgbm, pytorch, keras, nnetsauce in probabilistic Machine Learning (for longitudinal data) Reserving (work in progress) Jun 6, 2025
- R version of Probabilistic Machine Learning (for longitudinal data) Reserving (work in progress) Jun 5, 2025
- Probabilistic Machine Learning (for longitudinal data) Reserving (work in progress) Jun 4, 2025
- Python version of Beyond ARMA-GARCH: leveraging model-agnostic Quasi-Randomized networks and conformal prediction for nonparametric probabilistic stock forecasting (ML-ARCH) Jun 3, 2025
- Beyond ARMA-GARCH: leveraging model-agnostic Machine Learning and conformal prediction for nonparametric probabilistic stock forecasting (ML-ARCH) Jun 2, 2025
- Permutations and SHAPley values for feature importance in techtonique dot net's API (with R + Python + the command line) Jun 1, 2025
- Which patient is going to survive longer? Another guide to using techtonique dot net's API (with R + Python + the command line) for survival analysis May 31, 2025
- A Guide to Using techtonique.net's API and rush for simulating and plotting Stochastic Scenarios May 30, 2025
- Simulating Stochastic Scenarios with Diffusion Models: A Guide to Using techtonique.net's API for the purpose May 29, 2025
- Will my apartment in 5th avenue be overpriced or not? Harnessing the power of www.techtonique.net (+ xgboost, lightgbm, catboost) to find out May 28, 2025
- How long must I wait until something happens: A Comprehensive Guide to Survival Analysis via an API May 27, 2025
- Harnessing the Power of techtonique.net: A Comprehensive Guide to Machine Learning Classification via an API May 26, 2025
- Quantile regression with any regressor -- Examples with RandomForestRegressor, RidgeCV, KNeighborsRegressor May 20, 2025
- Survival stacking: survival analysis translated as supervised classification in R and Python May 5, 2025
- 'Bayesian' optimization of hyperparameters in a R machine learning model using the bayesianrvfl package Apr 25, 2025
- A lightweight interface to scikit-learn in R: Bayesian and Conformal prediction Apr 21, 2025
- A lightweight interface to scikit-learn in R Pt.2: probabilistic time series forecasting in conjunction with ahead::dynrmf Apr 20, 2025
- Extending the Theta forecasting method to GLMs, GAMs, GLMBOOST and attention: benchmarking on Tourism, M1, M3 and M4 competition data sets (28000 series) Apr 14, 2025
- Extending the Theta forecasting method to GLMs and attention Apr 8, 2025
- Nonlinear conformalized Generalized Linear Models (GLMs) with R package 'rvfl' (and other models) Mar 31, 2025
- Probabilistic Time Series Forecasting (predictive simulations) in Microsoft Excel using Python, xlwings lite and www.techtonique.net Mar 28, 2025
- Conformalize (improved prediction intervals and simulations) any R Machine Learning model with misc::conformalize Mar 25, 2025
- My poster for the 18th FINANCIAL RISKS INTERNATIONAL FORUM by Institut Louis Bachelier/Fondation du Risque/Europlace Institute of Finance Mar 19, 2025
- Interpretable probabilistic kernel ridge regression using Matérn 3/2 kernels Mar 16, 2025
- (News from) Probabilistic Forecasting of univariate and multivariate Time Series using Quasi-Randomized Neural Networks (Ridge2) and Conformal Prediction Mar 9, 2025
- Word-Online: re-creating Karpathy's char-RNN (with supervised linear online learning of word embeddings) for text completion Mar 8, 2025
- CRAN-like repository for most recent releases of Techtonique's R packages Mar 2, 2025
- Presenting 'Online Probabilistic Estimation of Carbon Beta and Carbon Shapley Values for Financial and Climate Risk' at Institut Louis Bachelier Feb 27, 2025
- Web app with DeepSeek R1 and Hugging Face API for chatting Feb 23, 2025
- tisthemachinelearner: A Lightweight interface to scikit-learn with 2 classes, Classifier and Regressor (in Python and R) Feb 17, 2025
- R version of survivalist: Probabilistic model-agnostic survival analysis using scikit-learn, xgboost, lightgbm (and conformal prediction) Feb 12, 2025
- Model-agnostic global Survival Prediction of Patients with Myeloid Leukemia in QRT/Gustave Roussy Challenge (challengedata.ens.fr): Python's survivalist Quickstart Feb 10, 2025
- A simple test of the martingale hypothesis in esgtoolkit Feb 3, 2025
- Command Line Interface (CLI) for techtonique.net's API Jan 31, 2025
- Gradient-Boosting and Boostrap aggregating anything (alert: high performance): Part5, easier install and Rust backend Jan 27, 2025
- Just got a paper on conformal prediction REJECTED by International Journal of Forecasting despite evidence on 30,000 time series (and more). What's going on? Part2: 1311 time series from the Tourism competition Jan 20, 2025
- Techtonique is released! (with a tutorial in various programming languages and formats) Jan 14, 2025
- Univariate and Multivariate Probabilistic Forecasting with nnetsauce and TabPFN Jan 14, 2025
- Just got a paper on conformal prediction REJECTED by International Journal of Forecasting despite evidence on 30,000 time series (and more). What's going on? Jan 5, 2025
- Python and Interactive dashboard version of Stock price forecasting with Deep Learning: throwing power at the problem (and why it won't make you rich) Dec 31, 2024
- Stock price forecasting with Deep Learning: throwing power at the problem (and why it won't make you rich) Dec 29, 2024
- No-code Machine Learning Cross-validation and Interpretability in techtonique.net Dec 23, 2024
- survivalist: Probabilistic model-agnostic survival analysis using scikit-learn, glmnet, xgboost, lightgbm, pytorch, keras, nnetsauce and mlsauce Dec 15, 2024
- Model-agnostic 'Bayesian' optimization (for hyperparameter tuning) using conformalized surrogates in GPopt Dec 9, 2024
- You can beat Forecasting LLMs (Large Language Models a.k.a foundation models) with nnetsauce.MTS Pt.2: Generic Gradient Boosting Dec 1, 2024
- You can beat Forecasting LLMs (Large Language Models a.k.a foundation models) with nnetsauce.MTS Nov 24, 2024
- Unified interface and conformal prediction (calibrated prediction intervals) for R package forecast (and 'affiliates') Nov 23, 2024
- GLMNet in Python: Generalized Linear Models Nov 18, 2024
- Gradient-Boosting anything (alert: high performance): Part4, Time series forecasting Nov 10, 2024
- Predictive scenarios simulation in R, Python and Excel using Techtonique API Nov 3, 2024
- Chat with your tabular data in www.techtonique.net Oct 30, 2024
- Gradient-Boosting anything (alert: high performance): Part3, Histogram-based boosting Oct 28, 2024
- R editor and SQL console (in addition to Python editors) in www.techtonique.net Oct 21, 2024
- R and Python consoles + JupyterLite in www.techtonique.net Oct 15, 2024
- Gradient-Boosting anything (alert: high performance): Part2, R version Oct 14, 2024
- Gradient-Boosting anything (alert: high performance) Oct 6, 2024
- Benchmarking 30 statistical/Machine Learning models on the VN1 Forecasting -- Accuracy challenge Oct 4, 2024
- Automated random variable distribution inference using Kullback-Leibler divergence and simulating best-fitting distribution Oct 2, 2024
- Forecasting in Excel using Techtonique's Machine Learning APIs under the hood Sep 30, 2024
- Techtonique web app for data-driven decisions using Mathematics, Statistics, Machine Learning, and Data Visualization Sep 25, 2024
- Parallel for loops (Map or Reduce) + New versions of nnetsauce and ahead Sep 16, 2024
- Adaptive (online/streaming) learning with uncertainty quantification using Polyak averaging in learningmachine Sep 10, 2024
- New versions of nnetsauce and ahead Sep 9, 2024
- Prediction sets and prediction intervals for conformalized Auto XGBoost, Auto LightGBM, Auto CatBoost, Auto GradientBoosting Sep 2, 2024
- Quick/automated R package development workflow (assuming you're using macOS or Linux) Part2 Aug 30, 2024
- R package development workflow (assuming you're using macOS or Linux) Aug 27, 2024
- A new method for deriving a nonparametric confidence interval for the mean Aug 26, 2024
- Conformalized adaptive (online/streaming) learning using learningmachine in Python and R Aug 19, 2024
- Bayesian (nonlinear) adaptive learning Aug 12, 2024
- Auto XGBoost, Auto LightGBM, Auto CatBoost, Auto GradientBoosting Aug 5, 2024
- Copulas for uncertainty quantification in time series forecasting Jul 28, 2024
- Forecasting uncertainty: sequential split conformal prediction + Block bootstrap (web app) Jul 22, 2024
- learningmachine for Python (new version) Jul 15, 2024
- learningmachine v2.0.0: Machine Learning with explanations and uncertainty quantification Jul 8, 2024
- My presentation at ISF 2024 conference (slides with nnetsauce probabilistic forecasting news) Jul 3, 2024
- 10 uncertainty quantification methods in nnetsauce forecasting Jul 1, 2024
- Forecasting with XGBoost embedded in Quasi-Randomized Neural Networks Jun 24, 2024
- Forecasting Monthly Airline Passenger Numbers with Quasi-Randomized Neural Networks Jun 17, 2024
- Automated hyperparameter tuning using any conformalized surrogate Jun 9, 2024
- Recognizing handwritten digits with Ridge2Classifier Jun 3, 2024
- Forecasting the Economy May 27, 2024
- A detailed introduction to Deep Quasi-Randomized 'neural' networks May 19, 2024
- Probability of receiving a loan; using learningmachine May 12, 2024
- mlsauce's `v0.18.2`: various examples and benchmarks with dimension reduction May 6, 2024
- mlsauce's `v0.17.0`: boosting with Elastic Net, polynomials and heterogeneity in explanatory variables Apr 29, 2024
- mlsauce's `v0.13.0`: taking into account inputs heterogeneity through clustering Apr 21, 2024
- mlsauce's `v0.12.0`: prediction intervals for LSBoostRegressor Apr 15, 2024
- Conformalized predictive simulations for univariate time series on more than 250 data sets Apr 7, 2024
- learningmachine v1.1.2: for Python Apr 1, 2024
- learningmachine v1.0.0: prediction intervals around the probability of the event 'a tumor being malignant' Mar 25, 2024
- Bayesian inference and conformal prediction (prediction intervals) in nnetsauce v0.18.1 Mar 18, 2024
- Multiple examples of Machine Learning forecasting with ahead Mar 11, 2024
- rtopy (v0.1.1): calling R functions in Python Mar 4, 2024
- ahead forecasting (v0.10.0): fast time series model calibration and Python plots Feb 26, 2024
- A plethora of datasets at your fingertips Part3: how many times do couples cheat on each other? Feb 19, 2024
- nnetsauce's introduction as of 2024-02-11 (new version 0.17.0) Feb 11, 2024
- Tuning Machine Learning models with GPopt's new version Part 2 Feb 5, 2024
- Tuning Machine Learning models with GPopt's new version Jan 29, 2024
- Subsampling continuous and discrete response variables Jan 22, 2024
- DeepMTS, a Deep Learning Model for Multivariate Time Series Jan 15, 2024
- A classifier that's very accurate (and deep) Pt.2: there are > 90 classifiers in nnetsauce Jan 8, 2024
- learningmachine: prediction intervals for conformalized Kernel ridge regression and Random Forest Jan 1, 2024
- A plethora of datasets at your fingertips Part2: how many times do couples cheat on each other? Descriptive analytics, interpretability and prediction intervals using conformal prediction Dec 25, 2023
- Diffusion models in Python with esgtoolkit (Part2) Dec 18, 2023
- Diffusion models in Python with esgtoolkit Dec 11, 2023
- Julia packaging at the command line Dec 4, 2023
- Quasi-randomized nnetworks in Julia, Python and R Nov 27, 2023
- A plethora of datasets at your fingertips Nov 20, 2023
- A classifier that's very accurate (and deep) Nov 12, 2023
- mlsauce version 0.8.10: Statistical/Machine Learning with Python and R Nov 5, 2023
- AutoML in nnetsauce (randomized and quasi-randomized nnetworks) Pt.2: multivariate time series forecasting Oct 29, 2023
- AutoML in nnetsauce (randomized and quasi-randomized nnetworks) Oct 22, 2023
- Version v0.14.0 of nnetsauce for R and Python Oct 16, 2023
- A diffusion model: G2++ Oct 9, 2023
- Diffusion models in ESGtoolkit + announcements Oct 2, 2023
- An infinity of time series forecasting models in nnetsauce (Part 2 with uncertainty quantification) Sep 25, 2023
- (News from) forecasting in Python with ahead (progress bars and plots) Sep 18, 2023
- Forecasting in Python with ahead Sep 11, 2023
- Risk-neutralize simulations Sep 4, 2023
- Comparing cross-validation results using crossval_ml and boxplots Aug 27, 2023
- Reminder Apr 30, 2023
- Did you ask ChatGPT about who you are? Apr 16, 2023
- A new version of nnetsauce (randomized and quasi-randomized 'neural' networks) Apr 2, 2023
- Simple interfaces to the forecasting API Nov 23, 2022
- A web application for forecasting in Python, R, Ruby, C#, JavaScript, PHP, Go, Rust, Java, MATLAB, etc. Nov 2, 2022
- Prediction intervals (not only) for Boosted Configuration Networks in Python Oct 5, 2022
- Boosted Configuration (neural) Networks Pt. 2 Sep 3, 2022
- Boosted Configuration (_neural_) Networks for classification Jul 21, 2022
- A Machine Learning workflow using Techtonique Jun 6, 2022
- Super Mario Bros © in the browser using PyScript May 8, 2022
- News from ESGtoolkit, ycinterextra, and nnetsauce Apr 4, 2022
- Explaining a Keras _neural_ network predictions with the-teller Mar 11, 2022
- New version of nnetsauce -- various quasi-randomized networks Feb 12, 2022
- A dashboard illustrating bivariate time series forecasting with `ahead` Jan 14, 2022
- Hundreds of Statistical/Machine Learning models for univariate time series, using ahead, ranger, xgboost, and caret Dec 20, 2021
- Forecasting with `ahead` (Python version) Dec 13, 2021
- Tuning and interpreting LSBoost Nov 15, 2021
- Time series cross-validation using `crossvalidation` (Part 2) Nov 7, 2021
- Fast and scalable forecasting with ahead::ridge2f Oct 31, 2021
- Automatic Forecasting with `ahead::dynrmf` and Ridge regression Oct 22, 2021
- Forecasting with `ahead` Oct 15, 2021
- Classification using linear regression Sep 26, 2021
- `crossvalidation` and random search for calibrating support vector machines Aug 6, 2021
- parallel grid search cross-validation using `crossvalidation` Jul 31, 2021
- `crossvalidation` on R-universe, plus a classification example Jul 23, 2021
- Documentation and source code for GPopt, a package for Bayesian optimization Jul 2, 2021
- Hyperparameters tuning with GPopt Jun 11, 2021
- A forecasting tool (API) with examples in curl, R, Python May 28, 2021
- Bayesian Optimization with GPopt Part 2 (save and resume) Apr 30, 2021
- Bayesian Optimization with GPopt Apr 16, 2021
- Compatibility of nnetsauce and mlsauce with scikit-learn Mar 26, 2021
- Explaining xgboost predictions with the teller Mar 12, 2021
- An infinity of time series models in nnetsauce Mar 6, 2021
- New activation functions in mlsauce's LSBoost Feb 12, 2021
- 2020 recap, Gradient Boosting, Generalized Linear Models, AdaOpt with nnetsauce and mlsauce Dec 29, 2020
- A deeper learning architecture in nnetsauce Dec 18, 2020
- Classify penguins with nnetsauce's MultitaskClassifier Dec 11, 2020
- Bayesian forecasting for uni/multivariate time series Dec 4, 2020
- Generalized nonlinear models in nnetsauce Nov 28, 2020
- Boosting nonlinear penalized least squares Nov 21, 2020
- Statistical/Machine Learning explainability using Kernel Ridge Regression surrogates Nov 6, 2020
- NEWS Oct 30, 2020
- A glimpse into my PhD journey Oct 23, 2020
- Submitting R package to CRAN Oct 16, 2020
- Simulation of dependent variables in ESGtoolkit Oct 9, 2020
- Forecasting lung disease progression Oct 2, 2020
- New nnetsauce Sep 25, 2020
- Technical documentation Sep 18, 2020
- A new version of nnetsauce, and a new Techtonique website Sep 11, 2020
- Back next week, and a few announcements Sep 4, 2020
- Explainable 'AI' using Gradient Boosted randomized networks Pt2 (the Lasso) Jul 31, 2020
- LSBoost: Explainable 'AI' using Gradient Boosted randomized networks (with examples in R and Python) Jul 24, 2020
- nnetsauce version 0.5.0, randomized neural networks on GPU Jul 17, 2020
- Maximizing your tip as a waiter (Part 2) Jul 10, 2020
- New version of mlsauce, with Gradient Boosted randomized networks and stump decision trees Jul 3, 2020
- Announcements Jun 26, 2020
- Parallel AdaOpt classification Jun 19, 2020
- Comments section and other news Jun 12, 2020
- Maximizing your tip as a waiter Jun 5, 2020
- AdaOpt classification on MNIST handwritten digits (without preprocessing) May 29, 2020
- AdaOpt (a probabilistic classifier based on a mix of multivariable optimization and nearest neighbors) for R May 22, 2020
- AdaOpt May 15, 2020
- Custom errors for cross-validation using crossval::crossval_ml May 8, 2020
- Documentation+Pypi for the `teller`, a model-agnostic tool for Machine Learning explainability May 1, 2020
- Encoding your categorical variables based on the response variable and correlations Apr 24, 2020
- Linear model, xgboost and randomForest cross-validation using crossval::crossval_ml Apr 17, 2020
- Grid search cross-validation using crossval Apr 10, 2020
- Documentation for the querier, a query language for Data Frames Apr 3, 2020
- Time series cross-validation using crossval Mar 27, 2020
- On model specification, identification, degrees of freedom and regularization Mar 20, 2020
- Import data into the querier (now on Pypi), a query language for Data Frames Mar 13, 2020
- R notebooks for nnetsauce Mar 6, 2020
- Version 0.4.0 of nnetsauce, with fruits and breast cancer classification Feb 28, 2020
- Create a specific feed in your Jekyll blog Feb 21, 2020
- Git/Github for contributing to package development Feb 14, 2020
- Feedback forms for contributing Feb 7, 2020
- nnetsauce for R Jan 31, 2020
- A new version of nnetsauce (v0.3.1) Jan 24, 2020
- ESGtoolkit, a tool for Monte Carlo simulation (v0.2.0) Jan 17, 2020
- Search bar, new year 2020 Jan 10, 2020
- 2019 Recap, the nnetsauce, the teller and the querier Dec 20, 2019
- Understanding model interactions with the `teller` Dec 13, 2019
- Using the `teller` on a classifier Dec 6, 2019
- Benchmarking the querier's verbs Nov 29, 2019
- Composing the querier's verbs for data wrangling Nov 22, 2019
- Comparing and explaining model predictions with the teller Nov 15, 2019
- Tests for the significance of marginal effects in the teller Nov 8, 2019
- Introducing the teller Nov 1, 2019
- Introducing the querier Oct 25, 2019
- Prediction intervals for nnetsauce models Oct 18, 2019
- Using R in Python for statistical learning/data science Oct 11, 2019
- Model calibration with `crossval` Oct 4, 2019
- Bagging in the nnetsauce Sep 25, 2019
- Adaboost learning with nnetsauce Sep 18, 2019
- Change in blog's presentation Sep 4, 2019
- nnetsauce on Pypi Jun 5, 2019
- More nnetsauce (examples of use) May 9, 2019
- nnetsauce Mar 13, 2019
- crossval Mar 13, 2019
- test Mar 10, 2019


Comments powered by Talkyard.